Table of Contents
Viruses that infect bacteria are known as phages or bacteriophages. Twort (19l5) described a degenerative alteration in staphylococcal colonies isolated from calf lymph that was transmissible serially via application of culture filtrates from the original growth. d’Herelle (1917) observed that the filtrates of dysentery patient faeces cultures produced transmissible lysis of a dysentery bacillus broth culture. He hypothesised that the lytic agent was a virus and designated it bacteriophage (phage: to eat). In nature, phages are commonly found in close interaction with bacteria. Phages are readily isolable from numerous habitats, including faeces, sewage, and other natural sources of mixed bacterial proliferation.
What are Bacteriophages?
- Bacteriophages are viruses of bacteria.
- They are the viruses that cause bacterial infection.
- They are obligate intracellular parasites that reproduce within bacteria by utilising some or all of the biosynthetic machinery of the host.
- Viruses are sometimes known as phages.
- Due to the presence of a nucleic acid genome encased in a protein coat, these extrachromosomal genetic components typically persist outside a host cell.
- Phages are prevalent in nature in intimate contact with bacteria and are widely disseminated in soil, excrement, and other environmental components.
- They are related with the transfer of genetic material between bacteria.
- Twort first reported the actions of bacteriophage in 1915, describing it as an infectious agent that altered the look of staphylococcal colonies.
- In 1979, d’Herelle proved the lytic activity of the culture filtrate on bacterial colonies. He hypothesised that the lytic agent was a virus and designated it bacteriophage (phage: to eat).
- The morphology of phages varies significantly. Also described are spherical or filamentous phages with DNA or RNA consisting of a single strand.
- The strong host specificity of viruses. They typically reside in the intestinal flora of humans and animals.
- They pass through screens that prevent bacteria from passing through.
- These are rendered inactive by boiling.
Role of Bacteriophages
- They play a key role in the translation of genetic information from one bacterium to another.
- In addition, they contribute to the evolution of bacterial kinds and the spread of certain virulence traits.
- In fact, phages may be helpful against bacterial infections, particularly those caused by antibiotic-resistant bacteria.
- Viruses have been utilised as cloning vectors in genetic engineering.
- They may have a role in bacterial population management in natural waterways.
Morphology of Bacteriophages
The majority of phages have a single, linear, double-stranded DNA genome. This genome is enveloped by a phage capsid protein coat. Typically, large phages consist of a head and a tail. The head encloses the genome, and the tail serves as both an attachment organ and a pathway for phage DNA to enter the host cell. The most thoroughly researched phages are the T-even phages (T2, T4, T6, etc.) that infect Escherichia coli. Traditionally, these phages serve as the model for characterising the morphology of bacteriophages. Phages have a hexagonal head and a cylindrical tail, like tadpoles.
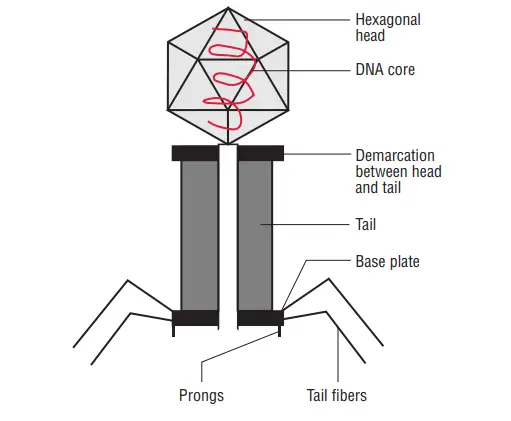
Head
- All phages possess a variable-sized and -shaped head structure.
- The form is icosahedral (20 sides) or filamentous.
- The length of the head ranges between 28 and 100 nanometers.
- In the majority of phages, the capsid of the bacteriophage head encloses a compartment containing the nucleic acid, which is a double-stranded DNA molecule.
- By virtue of its position within the bacteriophage head, this DNA is protected against destruction by environmental nucleases.
- However, a group of phages that infect solely male E. coli strains contains just RNA.
- The capsid or protein coat of a virus is formed of capsomeres, which are separate proteins.
Tail
- Many phages, although not all, have tails linked to their heads.
- During an infection, the nucleic acid flows via a hollow tube that comprises the tail.
- The tail of complex phages such as T4 is encircled by a contractile sheath and a terminal tail plate. The sheath contracts upon bacterial infection.
- The spikes and tail fibres come from the tail plate and bind particularly to receptors on the bacterial cell wall’s outer membrane.
Genome structure of Bacteriophage
Given that there are millions of different phages in the environment, phage genomes appear in numerous shapes and sizes. MS2 and other RNA phages have the smallest genomes, consisting of only a few kilobases. Nevertheless, some DNA phages, such as T4, may have large genomes containing hundreds of genes; the size and structure of the capsid varies with the genome size. The largest bacteriophage genomes measure 735 kilobytes.
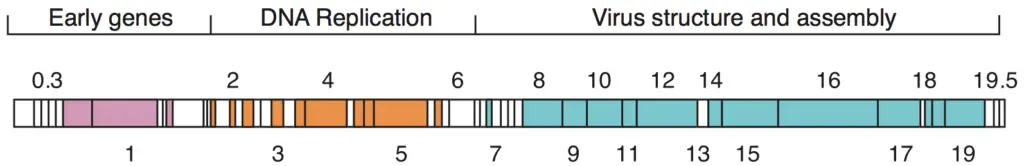
- The genomes of bacteriophages can be highly mosaic, meaning that the genomes of numerous phage species appear to be composed of numerous individual modules.
- These modules may exist in various configurations in other phage species. Mycobacteriophages, which are bacteriophages with mycobacterial hosts, are exceptional examples of this phenomenon.
- The genetic diversity of these mycobacteriophages may be the result of repeated instances of site-specific recombination and illegitimate recombination (the result of phage genome acquisition of bacterial host genetic sequences).
- The evolutionary mechanisms that shape the genomes of bacterial viruses vary among families and depend on the type of nucleic acid, the characteristics of the virion structure, and the viral life cycle.
- Some marine roseobacter phages’ genomic DNA contains deoxyuridine (dU) instead of deoxythymidine (dT).
- There is some evidence that this peculiar component is a means of evading bacterial defense mechanisms like restriction endonucleases and CRISPR/Cas systems, which evolved to recognize and cleave sequences within invading phages, thereby rendering them inactive.
- It has long been known that other phages utilize unusual nucleotides. Takahashi and Marmur discovered a Bacillus phage with dU replacing dT in its genome in 1963, and Kirnos and colleagues discovered a cyanophage containing 2-aminoadenine (Z) instead of adenine (A) in 1977.
Bacteriophage types Based on Replication
Virulent bacteriophages – Lytic cycle (cytoplasmic viral replication)
- Virulent bacteriophages are those that serve both our and their best interests. This variety of bacteriophage replicates via the lytic cycle.
- Lysis, also known as the lytic cycle, is a cytoplasmic viral replication process in which the bacteriophage injects its genetic material into a host cell, allowing this genetic material to replicate and produce numerous new phages.
- Once the host cell is full with newly formed bacteriophages, it ruptures from the inside, releasing the phages.
- It is essential to observe that all bacteriophages utilized for phage products and phage therapy are virulent.
Temperate bacteriophages – Lysogenic cycle
- Temperate bacteriophages are bacteriophages that replicate via the lysogenic cycle. In the lysogenc cycle, a phage infuses its generic material into a host, but instead of swiftly replicating, this generic material finds its way to the host’s genetic material and infuses itself with it, thereby becoming a prophage. It becomes part of the genetic material of the host, and when the host cell divides, the genetic material of the temperate phage replicates as well.
- Lysogenic bacteriophages continue to exist in this symbiotic state, in which they do not injure the host cell but rather use its resources invisibly. When a lysogenic bacteriophage senses that its survival is threatened, it can transition from the lysogenic cycle to the lytic cycle, resulting in the rapid replication of newly formed phages that burst out of the host cell.
- Bacteriophages that are capable of undergoing the lysogenic cycle, i.e., temperate phages, evaluate and select the appropriate cycle based on the environment in which they find themselves, i.e., lytic or lysogenic.
Bacteriophage Classification and Examples
Bacteriophages are categorized according to their nucleic acid content and morphology. There are 19 known families of bacteriophages, two of which are RNA bacteriophages. The primary families and characteristics of bacteriophages are described with examples in the following section.
| Examples | Family of the phage virus | Nucleic acid content | Morphological characteristics |
| T4(Escherichia virus T4) | Myoviridae | Linear dsDNA, 169 kbp long | Belongs to the Caudovirales order. These phages have elongated heads (approximately 110 nm in length), long tails with a collar (114 nm), spiked base plates, and six long tail fibers. They have a linear genome measuring between 33 kb and 244 kb and encoding between 40 and 415 proteins. These bacteriophages utilize cytoplasmic viral replication, are lytic, and lack lysogenic genes. |
| T2(Enterobacteria phage T2) | Myoviridae | Linear dsDNA, 170 kbp | Infects E.coli.The experiment performed by Hershey and Chase to demonstrate that DNA is a genetic material involved T2 phage infecting E. coli.The head is icosahedral and the tail is contractile.It passes through a lytic life cycle. |
| λ(Coliphage, Escherichia virus lambda) | Siphoviridae | Linear dsDNA, 48502 bp | Belongs to the Caudovirales order. Both bacteria and archaea are targets of these bacteriophages. However, many of the phages are known to target Lactobacillus, Streptococcus, Mycobacterium, and other bacteria. There are currently 313 species and 47 genera in this family. These phages lack an icosahedral enveloped head. Members are distinguished by their noncontractile, cross-banded tails. Their genomes are double-stranded, linear (approximately 50kb in size), and comprise 70 genes. These bacteriophages utilize cytoplasmic viral replication and leave the host cell via lysis. |
| M13(Escherichia virus M13) | Inoviridae | Circular ssDNA, 6407 nucleotides long | There is no assigned order for this virus family, which currently consists of 43 species divided into 2 genera. These bacteriophages specifically target microorganisms. These phages have a rod or filament geometry and lack an envelope. Their diameter is roughly 7nm and their length is roughly 2,000nm. Their genomes are circular, approximately 8 kb long, and code for four to ten proteins. These bacteriophages replicate via cytoplasmic means. |
| ΦX174 or phi x 174(Escherichia virus ΦX174) | Microviridae | Circular ssDNA, 5386 nucleotides | There is no assigned order for this virus family, which currently consists of 12 species divided among 7 genera. These bacteriophages specifically target Enterobacteria and Spiroplasma bacteria. These phages have round geometries and are icosahedral in shape. Their diameter is between 25 and 27 nanometers (each virion contains 60 copies of the F, G, and J proteins and 12 copies of the H protein). In addition, they have pentamers in the shape of trumpets, which are composed of five copies of the G protein and one copy of the H protein. Their DNA genome consists of a single strand. Although the majority of these bacteriophages have a lytic life cycle, a few have a temperate life cycle. The family name is derived from the ancient Greek word ” (mikrós), which means small, reflecting the fact that their genomes are among the tiniest of all DNA viruses. |
| Qβ, MS2, etc. | Leviviridae | ssRNA, 3500-4200 nucleotides | There is no assigned order for this virus family, which currently consists of four species divided into two genera. The bacteria that these bacteriophages target include Enterobacteria, Caulobacter, Pseudomonas, and Acinetobacter. These phages have spherical geometry and are icosahedral. Their diameter is roughly 26 nanometers. Their genomes are linear and unsegmented, approximately 4 kilobytes in size, and code for four proteins. To depart the host cell, these bacteriophages use the Positive stranded RNA virus transcription method and lysis. |
| Φ6, Φ7, Φ8, etc. | Cystoviridae | dsRNA, 14 kbp | This genus lacks an assigned order and contains only one species: Pseudomonas phage phi6. These bacteriophages target phaseolicola bacterial species. Their tripartite dsRNA genome, which is approximately 14 kb in length, and their protein and lipid outer layer are distinguishing characteristics. No other bacteriophage is known to possess an outer lipid layer. Currently, the majority of identified cystoviruses target Pseudomonas species. These phages have a spherical geometry and an icosahedral envelope. They are approximately 85nm in diameter. Their genomes are linear and segmented, with large genomes measuring 6,4 kb, medium genomes measuring 4 kb, and small genomes measuring 2.9 kb. The genome code encodes twelve proteins. These bacteriophages replicate via cytoplasmic means. |
International Committee for Taxonomy of Viruses (ICTV) classifies the bacteriophages under an order Caudovirales according to morphology and nucleic acid.
‘s the table:
| S.No. | Family | Morphology | Nucleic acid | Examples |
|---|---|---|---|---|
| 1 | Myoviridae | Non-enveloped, contractile tail | Linear dsDNA | T4 |
| 2 | Siphoviridae | Non-enveloped, long non-contractile tail | Linear dsDNA | Phage λ |
| 3 | Podoviridae | Non-enveloped, short non-contractile tail | Linear dsDNA | Coliphage T2 |
| 4 | Tectiviridae | Non-enveloped, isometric | Linear dsDNA | Phage PRD1 |
| 5 | Corticoviridae | Non-enveloped, isometric | Circular dsDNA | PM2 |
| 6 | Lipothrixviridae | Enveloped, rod-shaped | Linear dsDNA | α-lipothrixviruses, β-lipothrixviruses |
| 7 | Plasmaviridae | Enveloped, pleomorphic | Circular dsDNA | Acholeplasma phage, Sma phage |
| 8 | Rudiviridae | Non-enveloped, rod-shaped | Linear dsDNA | Rudivirus |
| 9 | Fuselloviridae | Non-enveloped, lemon-shaped | Circular dsDNA | SSV-1 |
| 10 | Inoviridae | Non-enveloped, filamentous | Circular ssDNA | Coliphage fd, MS2, φx174 |
| 11 | Microviridae | Non-enveloped, isometric | Circular dsDNA | Spiroplasma |
| 12 | Leviviridae | Non-enveloped, isometric | Linear ssRNA | Coliphage Qβ |
| 13 | Cystoviridae | Enveloped, spherical | Segmented dsRNA | φ6 |
On the bases of electron microscopic studies Bradley (1967) described seven morphological types:
| Type | Description | Nucleic acid | Examples |
|---|---|---|---|
| A | Hexagonal head, rigid tail with contractile sheath and tail fibers | dsDNA | T2, T4, T6 phage |
| B | Hexagonal head, lacks contractile sheath, tail flexible | dsDNA | T1, T5 phage |
| C | Hexagonal head, tail shorter than head, lacks contractile sheath | dsDNA | T3, T7 phage |
| D | Head made up of capsomers, lacks tail | ssDNA | φX174 phage |
| E | Head with small capsomers, no tail | ssRNA | F2, MS2 phage |
| F | Filamentous phage | ssDNA | Fd, F1 phage |
| G | Has a lipid-containing envelope, no detectable capsid | dsRNA | MV-L2 phage |
Life Cycle of Bacteriophages
Phages have two distinct life cycle types. The virulent or lytic cycle concludes with the lysis of the host bacterium and the release of progeny virions. During the temprate or lysogenic cycle, the phage DNA integrates into the bacterial genome and replicates synchronously with it, without harming the host cell.
Lytic Cycle
The steps of replication of a virulent phage are adsorption, penetration, synthesis of phage components, assembly, maturation, and release of offspring phage particles.
i. Adsorption
- The tail of phage particles attaches to virus-specific receptors on the host cell through random collision.
- Adsorption is a unique process dependent on the existence of complimentary chemical groups on the receptor sites on the bacterial surface and on the phage’s terminal base plate.
- Under perfect conditions, adsorption is a process that takes only minutes to complete.
- Any component on the bacterial surface can function as a phage receptor. Phage host specificity is determined at the adsorption level.
- Even bacterial strains that are resistant to infection by the complete phage can be experimentally infected by injecting phage DNA directly.
- Transfection is the infection of a bacteria by the nucleic acid of a naked phage.
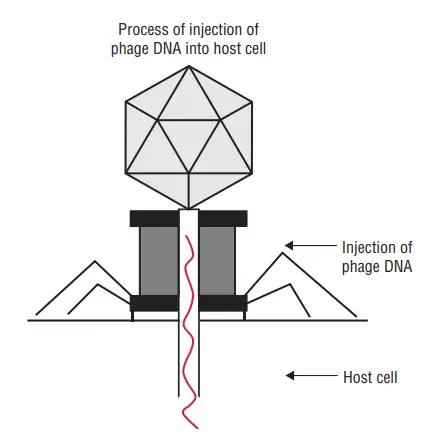
ii. Penetration
- Following adsorption, the majority of phages inject their nucleic acid into the bacterial cytoplasm while leaving their protein capsid outside.
- Similar to an injection with a syringe, penetration resembles injection. Six tail pins establish contact with the host cell surface and securely bind the phage plate to it upon adsorption.
- The contractile tail sheath then contracts, driving the tail tube’s hollow interior into the bacterial cell wall.
- The phage DNA then traverses the tail’s interior hollow tube.
- The phage’s empty head and tail stay outside the bacterium as the shell or ‘ghost’ following penetration.
- The presence of lysozyme on the phage tail, which creates a hole in the bacterial cell wall for the entry of the phage core, may promote bacterial penetration.
- When bacteria are mixed with phage particles at a high multiplicity (that is, a very large number of phages per bacterial cell), many holes are formed on the bacterial cell, resulting in the loss of cell contents. Without viral replication, lysis of bacteria occurs. This is referred to as external lysis.
iii. Synthesis of Phage Nucleic Acid and Proteins
- The manufacturing of phage components begins as soon as the phage nucleic acid penetrates the host cell.
- The earliest products to be generated (referred to as early proteins) are the enzymes required for the construction of the complex compounds that are unique to the phage.
- Subsequently, late proteins such as the protein subunits of the phage head and tail are produced.
- During this time, bacterial protein, DNA, and RNA synthesis ceases, and the cell is pushed to produce viral components.
- Late genes are expressed only after phage DNA replication has occurred.
- The structural components of new phage particles, including their heads, tails, and fibres, are encoded by late gene products.
- Phage lysozyme, which degrades the bacterial cell wall to release the mature phage particles, is also a late product.
iv. Assembly and Maturation
- In order to create the mature progeny phage particle, the structural proteins and nucleic acid of the phage assemble along certain paths.
- In the bacterial cell, phage DNA, head protein, and tail protein are generated independently.
- The DNA is compacted into a polyhedron and ‘packed’ into the head; the tail structures are then inserted.
- This process of assembling phage components into an infectious mature phage particle is known as maturation.
v. Release
- At the end of the intracellular phase, many phages lyse their host cells.
- Typically, the adult progeny phages are released by the lysis of the bacterial cell.
- During phage replication, the bacterial cell wall is weakened and develops a spherical form.
- The weakening cell wall is ruptured or lysed by phage enzymes, resulting in the release of mature daughter phages.
Eclipse Phase
- The time between the entry of the phage nucleic acid into the bacterial cell and the appearance of the first infectious intracellular phage particle is referred to as the eclipse phase.
- It is the time necessary for the synthesis of phage components and their assembly into mature phage particles.
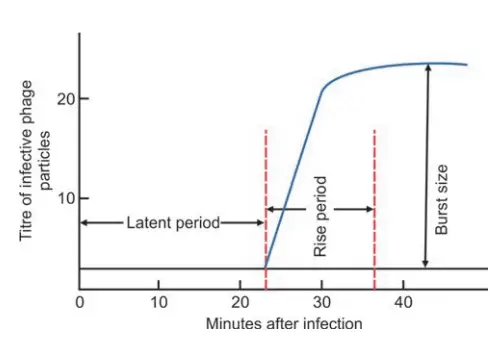
- Latent period refers to the time between the infection of a bacterial cell and the initial release of infectious phage particles (20 to 40 minutes).
- Immediately following the latent period, the number of phage particles released grows until the maximum number of daughter phages is reached.
- The period in which the quantity of infectious phages released increases is referred to as the rising phase.
- The average number of offspring phages produced per infected bacterial cell is referred to as the burst size (100 to 300 phages).
- Experiments are conducted in which one phage is used to infect one bacteria, and the discharge of infected phage particles is measured serially over a period of time.
- The plotted results of such an experiment are referred to as the one-step growth curve.
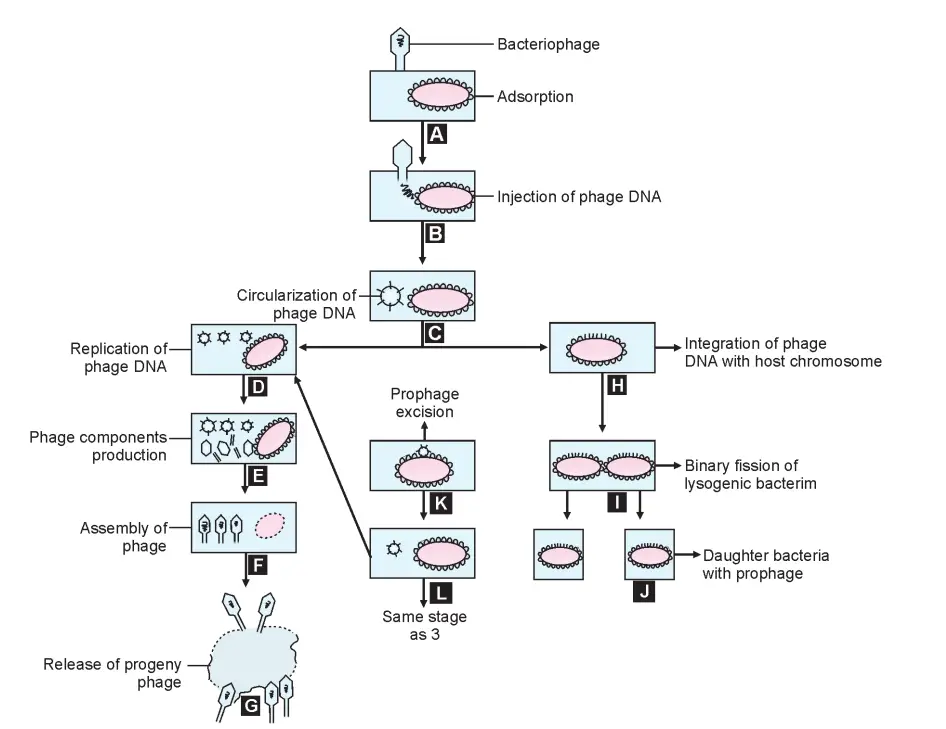
Lysogenic Cycle
- In contrast to virulent phages, which result in the lysis of the host cell, temperate phages form symbiotic relationships with their hosts without harming them.
- After entering the host cell, the nucleic acid of a temperate phage becomes incorporated into the bacterial chromosome.
- The nucleic acid of the integrated phage is known as the prophage.
- The prophage replicates synchronously with the host chromosome, mimicking its behaviour.
- This behaviour is known as lysogeny, and a bacterium that harbours a prophage within its genome is known as a lysogenic bacterium or lysogen, as the prophage retains the ability to lyse its host bacterium. Phages capable of entering into this relationship are known as temperate phages.
- In most cases, the prophage is integrated into the bacterial genome, but it can also live independently. The lysogenization process has little effect on bacterial metabolism.
- In some instances, temperate phages contribute to the pathogenicity of gram-positive and gram-negative bacteria isolated from clinical settings.
- The prophage imparts new characteristics to the lysogenic bacterium. This process is referred to as lysogenic or phage conversion.
- The existence of prophage b in Corynebacterium diphtheriae determines its toxin production. Elimination of this prophage eliminates the toxigenicity of the bacillus, whereas lysogenization can transform nontoxigenic strains into toxigenic ones.
- Types C and D of Clostridium botulinum only generate toxin when infected with phages CE b and DE b, respectively.
- Salmonella phages in temperate environments can alter the antigenic characteristics of somatic O antigen. When Salmonella is infected by an epsilon phage, the outer lipopolysaccharide layer may undergo structural changes. S. Anatum has an antigenic formula of 3, 10: e, h: 1,6; however, when lysogenized by a temperate phage, its antigenic formula changes to 3, 15: e, h: 1,6.
- During the proliferation of lysogenic bacteria, the prophage may become “excised” from some cells.
- The excised prophage commences lytic replication and releases daughter phage particles that infect and lysogenize additional bacterial cells. This is characterised as spontaneous prophage induction.
- Certain physical and chemical agents can encourage all lysogenic bacteria in a population to switch to the lytic cycle, although this is an uncommon occurrence.
- Such agents include ultraviolet light, hydrogen peroxide, and nitrogen mustard. A lysogenic bacterium is resistant to reinfection by identical or similar phages. The term for this is superinfection immunity.
Examples of phage conversions
The examples of phage conversions are as follows:
- Phage-mediated conversion of Salmonella somatic antigens: This occurs when a significant number of temperate Salmonella phages alter the antigenic characteristics of the somatic O antigen. When lysogenized by a temperate phage, Salmonella Anatum (antigenic formula: 3,10:e, h:1,6) reveals a new antigenic formula (3,15:e, h:1,6), which is an antigenic formula of Salmonella Newington.
- Corynebacterium diphtheriae’s toxicity: Toxin synthesis in C. diphtheriae is attributable to the presence of prophage beta (). By removing the prophage, the bacteria become nontoxigenic. Similarly, nontoxigenic C. diphtheriae can be rendered toxic via phage-mediated lysogenization.
- Toxicity of Clostridium botulinum: Clostridium botulinum toxicity is dictated by the presence of phages CE and DE in C. botulinum types C and D, respectively. Elimination of the phages eradicates the bacillus’s toxigenicity.
- Transduction: Bacteriophages can transport genes from one bacterium to another via transduction. This is referred to as transduction. This behaviour has been observed in numerous bacterial species and is considered as one of the primary routes by which genetic material is transferred between bacteria in nature.
Significance of Phages
1. Virulent Phage
i. Phage Assay
- When a phage is given to a sensitive bacterial lawn culture, clearance occurs following incubation. These lysis zones are known as plaques.
- The size, shape, and composition of plaques are unique to each phage. Plaque assay can be used to determine the amount of viable phages in a sample since, under optimal conditions, a single phage particle can produce one plaque.
- Plaquing is also useful for purifying phages since plaques are similar to bacterial colonies.
ii. Phage Typing
- Typing of bacteriophages (phages) is determined by the specificity of phage surface receptors for cell surface receptors.
- Many phages can be employed as epidemiological markers to distinguish between biochemically or serologically indistinguishable bacterial strains due to their restricted host range.
- Staphylococcus aureus, Salmonella typhi, and Vibrio cholerae outbreaks have been traced using this technique.
- The to-be-typed strain is inoculated onto a nutrient agar plate to form a grass culture. After phages have been dried, a predetermined dose is administered to squares that have been designated (routine test dose).
- The maximum dilution of the phage preparation that achieves confluent lysis is referred to as the standard test dose (RTD).
- Some phages will lyse the culture after an overnight incubation, whereas others will not.
- The designation of the phage(s) that lyse the strain expresses the phage type of the strain. Plaque refers to the area of lysis caused by a phage.
- Since a single phage particle is capable of forming a single plaque, the plaque assay can be used to quantify the amount of viable phages in a sample.
- The most crucial use of phage typing is intraspecies typing of bacteria, such as the phage typing of S. typhi and staphylococci.
- There are phages that lyse all members of a bacterial genus (e.g., Salmonella), all members of a species (e.g., B. anthracis), and all members of a biotype or subspecies (e.g., Mukerjee’s phage IV, which lyses all strains of classical V. cholerae but not V. cholerae biotype EI Tor).
2. Temperate Phage
i. Transduction
- Bacteriophages can operate as gene carriers from one bacterium to the next. This is referred to as transduction.
- There are two recognised types of transduction: generalised transduction, in which any piece of the donor DNA can be transmitted, and specialised transduction, in which only a particular set of genes can be transported to a recipient cell.
ii. Toxin Production
- The synthesis of toxins by Corynebacterium diphtheriae and Clostridium botulinum types C and D is determined by prophage DNA-carrying genes.
iii. Antigenic Property
- Salmonella phages in temperate environments can alter the antigenic characteristics of somatic O antigen.
- This acquisition of new capabilities by infected bacterial cells is known as “phage conversion.”
iv. Cloning Vector
- In genetic modifications, bacteriophages have been utilised as cloning vectors.
Lytic Cycle vs Lysogenic Cycle
| Characteristics | Lytic Cycle | Lysogenic Cycle |
|---|---|---|
| Definition | Immediate replication of viral genome and destruction of host cell | Integration of viral genome into host cell’s genome without immediate destruction of host cell |
| Host cell damage | Leads to destruction of host cell | Does not lead to destruction of host cell |
| Viral replication | Immediate replication of viral genome after infecting host cell | Integration of viral genome into host cell’s genome and replication during cell division |
| Viral gene expression | Viral genes expressed immediately leading to production of viral proteins | Viral genes may remain dormant and not expressed for some time |
| Prophage formation | Does not occur | Viral genome integrates into host cell’s genome and forms a prophage |
| Excision of prophage | Does not apply | Prophage can be excised from host cell’s genome and enter the lytic cycle |
| Host range | Narrow | Broad |
| Efficiency of infection | High | Low |
| Multiplication rate | Rapid | Slow |
| Number of virions produced | High | Low |
| Evolutionary advantage | Rapidly spreads virus and allows for more virions to be produced | Allows virus to coexist with host cell and may confer benefits to host |
| Examples of phages | T4, T7 | Lambda, Mu |
| Trigger for cycle switch | N/A | Environmental stress or damage to host cell DNA |
| Medical significance | Potential use in phage therapy | Role in pathogenicity and gene transfer in bacteria |
Applications of Bacteriophages
Bacteriophages, or simply phages, have numerous applications in various fields. Here are some of them:
- Phage therapy: Phages have been used to treat bacterial infections in humans and animals, particularly in cases where antibiotic resistance is a concern.
- Food industry: Phages are used as a natural method to control bacterial pathogens in food products. They can be used to control foodborne pathogens like Salmonella, Listeria, and E. coli.
- Agriculture: Phages can be used to control bacterial plant diseases, particularly in crops like fruits and vegetables.
- Biotechnology: Phages are useful tools in biotechnology, particularly in genetic engineering and bioremediation.
- Biocontrol of pests: Phages can be used to control pests like mosquitoes and other insects that carry harmful diseases.
- Biosensors: Phages can be used as biosensors to detect bacterial contaminants in food and water.
- Research: Phages are important tools in genetic research, particularly in the study of bacterial genetics and gene transfer.
- Antibiotic development: Phages can be used to screen and identify new antibiotics that are effective against bacterial infections.
What is Phage Therapy?
- Phage therapy is a type of medical treatment that uses bacteriophages, or simply phages, to target and kill specific bacteria that cause infections. Phages are viruses that infect and replicate within bacteria, ultimately leading to their destruction.
- In phage therapy, specific phages that are effective against the target bacteria are identified and isolated. These phages are then administered to the patient, either orally or topically, to treat the infection. The phages infect and kill the bacteria, ultimately reducing the symptoms of the infection and promoting healing.
- Phage therapy is particularly useful in cases where bacterial infections are resistant to antibiotics. This is because phages are highly specific in their targeting of bacteria, and can be tailored to target specific strains of bacteria. Phage therapy also has fewer side effects than traditional antibiotic treatments, since phages target only the specific bacteria causing the infection, leaving the healthy bacteria in the body untouched.
- However, phage therapy is still an emerging field, and more research is needed to fully understand its potential benefits and limitations.
Phage Therapy Principle
- The principle of phage therapy is to use bacteriophages, or simply phages, to target and kill specific bacteria that cause infections. Phages are viruses that infect and replicate within bacteria, ultimately leading to their destruction.
- Phage therapy involves identifying and isolating specific phages that are effective against the target bacteria. These phages are then administered to the patient, either orally or topically, to treat the infection. The phages infect and kill the bacteria, ultimately reducing the symptoms of the infection and promoting healing.
- Phage therapy has several advantages over traditional antibiotic treatments. Phages are highly specific in their targeting of bacteria, and can be tailored to target specific strains of bacteria. This means that phage therapy is particularly useful in cases where bacterial infections are resistant to antibiotics. Phage therapy also has fewer side effects than traditional antibiotic treatments, since phages target only the specific bacteria causing the infection, leaving the healthy bacteria in the body untouched.
- However, there are also some limitations to phage therapy. One of the main limitations is the potential for the bacteria to develop resistance to the phages over time. Additionally, the use of phage therapy requires identifying and isolating the specific phages that are effective against the target bacteria, which can be a time-consuming process. Despite these limitations, phage therapy is an emerging field with promising potential for treating bacterial infections that are resistant to traditional antibiotics.
Advantages of Phage Therapy
Phage therapy has several advantages over traditional antibiotic treatments:
- Specificity: Phages are highly specific in their targeting of bacteria, and can be tailored to target specific strains of bacteria. This means that phage therapy is particularly useful in cases where bacterial infections are resistant to antibiotics.
- Safety: Phages are naturally occurring viruses that are typically harmless to humans, and have fewer side effects than traditional antibiotic treatments. This is because phages target only the specific bacteria causing the infection, leaving the healthy bacteria in the body untouched.
- Versatility: Phages can be used to treat a wide range of bacterial infections, including those that are difficult to treat with traditional antibiotics.
- No drug resistance: Unlike antibiotics, phages have the ability to co-evolve with bacteria, preventing them from developing resistance over time.
- Rapid action: Phages have the ability to replicate rapidly within the host, leading to a rapid and effective response against bacterial infections.
- Easy to administer: Phages can be administered orally, topically or by injection, making them easy to use in a variety of clinical settings.
- Cost-effective: Phage therapy is often less expensive than traditional antibiotic treatments, as phages can be easily produced and do not require complex manufacturing processes.
Limitations of Phage Therapy
While phage therapy has many potential benefits, there are also some limitations and challenges associated with this approach:
- Limited knowledge: Despite decades of research on phages, there is still much we don’t know about their interactions with bacteria and the human body.
- Narrow spectrum of activity: Phages are highly specific in their targeting of bacteria, and may not be effective against all strains of a particular bacterial species. This can limit their use in some cases.
- Emergence of resistance: Although phages can co-evolve with bacteria, there is still a risk that bacterial strains will eventually develop resistance to the phages.
- Production challenges: The production of phages can be challenging, particularly when large quantities are needed for clinical use.
- Regulatory issues: The use of phage therapy is not yet widely regulated, and there are few guidelines in place for its clinical use.
- Safety concerns: Although phages are generally safe, there is still a risk of adverse effects, particularly in people with weakened immune systems.
- Lack of clinical trials: While there have been many laboratory studies on phage therapy, there is still a lack of large-scale clinical trials to demonstrate its safety and efficacy.
Phage Therapy vs Antibiotics
| Phage Therapy | Antibiotics |
|---|---|
| Targeted | Broad-spectrum |
| Highly specific to bacterial strains | May target both harmful and beneficial bacteria |
| Can co-evolve with bacteria, reducing the risk of resistance | Resistance can develop quickly |
| No known side effects | May have side effects, including allergic reactions and disruption of gut microbiome |
| Generally safe for use in people with weakened immune systems | May not be safe for use in people with weakened immune systems |
| Production can be challenging, but can be targeted to specific bacterial strains | Production is well-established and can be done on a large scale |
| Limited regulatory guidelines | Well-established regulatory guidelines |
| Less research and fewer clinical trials, but growing interest | Extensive research and many clinical trials |
| Cost-effective in some cases | Generally more expensive |
| May take longer to see results | Results are typically faster |
What is Phage typing?
- Phage typing is a laboratory technique used to identify and differentiate bacterial strains based on their susceptibility to various bacteriophages.
- It involves exposing a bacterial isolate to a panel of different bacteriophages and observing which phages are able to infect and lyse the bacteria.
- By comparing the phage susceptibility patterns of different bacterial strains, researchers can identify similarities and differences between them and use this information to classify and trace the origin of bacterial isolates.
- Phage typing can be particularly useful in outbreak investigations and in tracking the spread of bacterial infections within a community or population.
Limitations or Challenges of Bacteriophages
While bacteriophages offer several potential advantages as therapeutic agents, there are also some limitations and challenges associated with their use:
- Limited host range: Each bacteriophage is specific to certain bacterial species or strains, which can make it difficult to find a phage that is effective against a particular bacterial infection.
- Emergence of resistant bacteria: As with antibiotics, the overuse or misuse of bacteriophages can lead to the emergence of resistant bacterial strains that are no longer susceptible to phage therapy.
- Limited stability: Bacteriophages can be sensitive to environmental factors such as pH, temperature, and UV light, which can limit their shelf life and effectiveness.
- Safety concerns: While bacteriophages are generally considered safe, there is still a need for more rigorous safety testing to ensure that they do not have harmful effects on the human body.
- Regulatory challenges: The regulatory pathways for approving bacteriophage-based therapies are still unclear, which can make it difficult for researchers and companies to develop and bring phage-based products to market.
- Lack of standardized methods: There is currently no standardized method for evaluating the efficacy of phage therapy, which can make it difficult to compare results across different studies.
- Cost: Developing and manufacturing bacteriophage-based therapies can be expensive, which may limit their accessibility to patients in need.
FAQ
What are bacteriophages and how do they work?
Bacteriophages, or phages for short, are viruses that infect and kill bacteria. They work by attaching to specific receptors on the surface of bacterial cells and injecting their genetic material (DNA or RNA) into the cell. Once inside, the phage uses the host cell’s machinery to replicate itself, eventually causing the cell to burst open and release new phages that can infect other bacteria.
What is the difference between the lytic and lysogenic cycles of bacteriophages?
The lytic cycle involves the immediate replication of the phage inside the host cell, eventually leading to its destruction and release of new phages. The lysogenic cycle, on the other hand, involves the integration of the phage DNA into the host cell’s genome, where it can remain dormant for long periods of time. During this cycle, the host cell replicates normally and passes on the phage DNA to its daughter cells.
Can bacteriophages be used to treat bacterial infections in humans?
Yes, bacteriophages can be used to treat bacterial infections in humans. This is known as phage therapy and involves the use of specific phages to target and kill the bacteria causing the infection. Phage therapy is still in the experimental stages and is not widely used in clinical practice.
How specific are bacteriophages in targeting certain bacterial species or strains?
Bacteriophages are highly specific in their targeting of certain bacterial species or strains. Each phage is designed to recognize and attach to a specific receptor on the surface of the bacterial cell, meaning that it will only infect and replicate within that particular species or strain of bacteria.
Can bacteria develop resistance to bacteriophages like they do with antibiotics?
Yes, bacteria can develop resistance to bacteriophages, although the mechanisms for doing so are different than those for developing resistance to antibiotics. Bacteria can modify or lose the receptors on their surface that the phage uses to attach and enter the cell, effectively making them immune to that particular phage.
Are bacteriophages safe for human use and consumption?
Bacteriophages are generally considered safe for human use and consumption, as they only target and kill bacteria and do not infect human cells. However, more research is needed to fully evaluate their safety and potential side effects.
How are bacteriophages produced and purified for use in research and therapy?
Bacteriophages can be isolated from natural sources, such as soil or water, or from bacterial cultures. Once isolated, they can be grown and propagated in bacterial hosts, purified using various techniques, and stored for later use in research or therapy.
Can bacteriophages be used in combination with antibiotics to enhance their efficacy?
Yes, bacteriophages can be used in combination with antibiotics to enhance their efficacy. This approach is known as phage-antibiotic synergy and involves using phages to target and weaken bacteria, making them more susceptible to antibiotics.
What are some of the challenges in developing and implementing bacteriophage therapy?
Some of the challenges in developing and implementing bacteriophage therapy include the specificity of phages, the potential for bacterial resistance, the need for careful selection and matching of phages to specific bacterial strains, and the lack of regulatory approval in many countries.
Are bacteriophages naturally occurring or can they be genetically engineered?
Bacteriophages are naturally occurring, but they can also be genetically engineered to enhance their targeting specificity or other properties. Genetic engineering techniques can be used to modify the phage genome, such as by deleting genes that encode for virulence factors or adding genes that increase phage stability or efficacy.
References
- Kasman LM, Porter LD. Bacteriophages. [Updated 2022 Sep 26]. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2023 Jan-. Available from: https://www.ncbi.nlm.nih.gov/books/NBK493185/
- Sulakvelidze A, Alavidze Z, Morris JG Jr. Bacteriophage therapy. Antimicrob Agents Chemother. 2001 Mar;45(3):649-59. doi: 10.1128/AAC.45.3.649-659.2001. PMID: 11181338; PMCID: PMC90351.
- Principi, N., Silvestri, E., & Esposito, S. (2019). Advantages and Limitations of Bacteriophages for the Treatment of Bacterial Infections. Frontiers in Pharmacology, 10. doi:10.3389/fphar.2019.00513
- Chen, Y., Batra, H., Dong, J., Chen, C., Rao, V. B., & Tao, P. (2019). Genetic Engineering of Bacteriophages Against Infectious Diseases. Frontiers in Microbiology, 10. doi:10.3389/fmicb.2019.00954
- Haq, I.U., Chaudhry, W.N., Akhtar, M.N. et al. Bacteriophages and their implications on future biotechnology: a review. Virol J 9, 9 (2012). https://doi.org/10.1186/1743-422X-9-9
- Moineau, S. (2013). Bacteriophage. Brenner’s Encyclopedia of Genetics, 280–283. doi:10.1016/b978-0-12-374984-0.00131-5
- Rifkind, D., & Freeman, G. L. (2005). BACTERIOPHAGE. The Nobel Prize Winning Discoveries in Infectious Diseases, 103–105. doi:10.1016/b978-012369353-2/50022-9
- Yata, T., Nianiaris, N., Songsivilai, S., & Hajitou, A. (2014). Bacteriophage. Gene Therapy of Cancer, 479–490. doi:10.1016/b978-0-12-394295-1.00033-0
- Kutter, E. (2001). Bacteriophages. Encyclopedia of Genetics, 179–186. doi:10.1006/rwgn.2001.0106
- Abedon, S. T., Duffy, S., & Turner, P. E. (2009). Bacteriophage Ecology. Encyclopedia of Microbiology, 42–57. doi:10.1016/b978-012373944-5.00022-5
- Rees, C. E. D., Swift, B. M. C., & Botsaris, G. (2014). Bacteriophage-Based Techniques for Detection of Foodborne Pathogens. Encyclopedia of Food Microbiology, 194–202. doi:10.1016/b978-0-12-384730-0.00032-x
- https://www.nature.com/scitable/definition/bacteriophage-phage-293/
- https://www.britannica.com/science/bacteriophage
- https://www.khanacademy.org/science/biology/biology-of-viruses/virus-biology/a/bacteriophages
