Table of Contents
The technique of homology modeling, which is also referred to as comparative modeling, is a robust computational approach that is widely employed in the domain of structural biology. The technique enables researchers to forecast the spatial arrangement of a protein or any other macromolecule by utilizing its amino acid sequence and the established configuration of a corresponding protein. Homology modeling is a useful technique that leverages the principle of structural and functional similarity among proteins with similar sequences. This approach offers valuable insights into the relationships between protein structure and function. The aforementioned methodology finds utility in diverse domains such as pharmaceutical exploration, protein manipulation, and comprehension of biological mechanisms. The utility of homology models is contingent upon the degree of sequence similarity exhibited between the target and template proteins. However, homology modeling has emerged as a valuable technique for investigating proteins in instances where experimental approaches are arduous or time-prohibitive.
What is Homology Modeling?
Homology modeling, also referred to as comparative modeling, is a computational technique utilized in structural biology to predict the three-dimensional (3D) structure of a protein or other macromolecule based on its amino acid sequence and the known structure of a related protein. It is especially useful when experimental techniques like X-ray crystallography or nuclear magnetic resonance (NMR) spectroscopy are impractical or time-consuming.
The underlying principle of homology modeling is that proteins with similar sequences frequently assume similar structures and functions. If the sequences of two proteins are highly similar, it is likely that their structures will also be similar. If the structure of one protein is known, it can be used as a model to determine the structure of another protein with a similar sequence.
Software for Homology Modeling
There are a variety of software programs available for homology modeling, each with its own set of advantages and capabilities. Here are some widely used homology modeling software programs:
- MODELLER: MODELLER is a well-known software application for comparative protein structure modeling. It combines spatial constraint satisfaction, genetic algorithm, and molecular dynamics to generate precise protein models. MODELLER offers a vast array of options for model construction, refinement, and evaluation.
- SWISS-MODEL: SWISS-MODEL is an automated server for protein homology modeling that provides user-friendly web interfaces. It uses a large database of template structures in conjunction with multiple algorithms and optimization techniques to generate high-quality models. Additionally, SWISS-MODEL provides additional instruments for model evaluation and visualization.
- Rosetta: Rosetta is a suite of software with modules for protein structure prediction and homology modeling. Utilizing energy functions and optimization algorithms, it explores conformational space and generates precise models. Rosetta permits users to modify modeling protocols and incorporate experimental data into the modeling procedure.
- I-TASSER: I-TASSER (Iterative Threading ASSembly Refinement) is a commonly used software for protein structure prediction, which includes both ab initio modeling and homology modeling. It uses threading-based algorithms to generate initial 3D models and then refines the models through iterative fragment assembly and refinement. I-TASSER provides comprehensive evaluation and visualization tools for structures.
- Phyre2: Phyre2 is an online application for modeling and analyzing proteins. For homology modeling, it employs a combination of profile-based and ab initio modeling techniques. Phyre2 offers user-friendly interfaces, automated workflows, and comprehensive model quality evaluation to facilitate the creation of reliable protein models.
- Modeller (UCSF Chimera): Modeller is an integral module of the widely used molecular visualization software UCSF Chimera. It enables users to perform homology modeling directly within the graphic user interface of Chimera. Chimera’s integration of modeling and visualization capabilities facilitates model creation, refinement, and analysis.
Homology Modeling Steps
The homology modeling procedure encompasses various crucial stages. Provided below is a comprehensive outline of the procedures entailed in homology modeling.
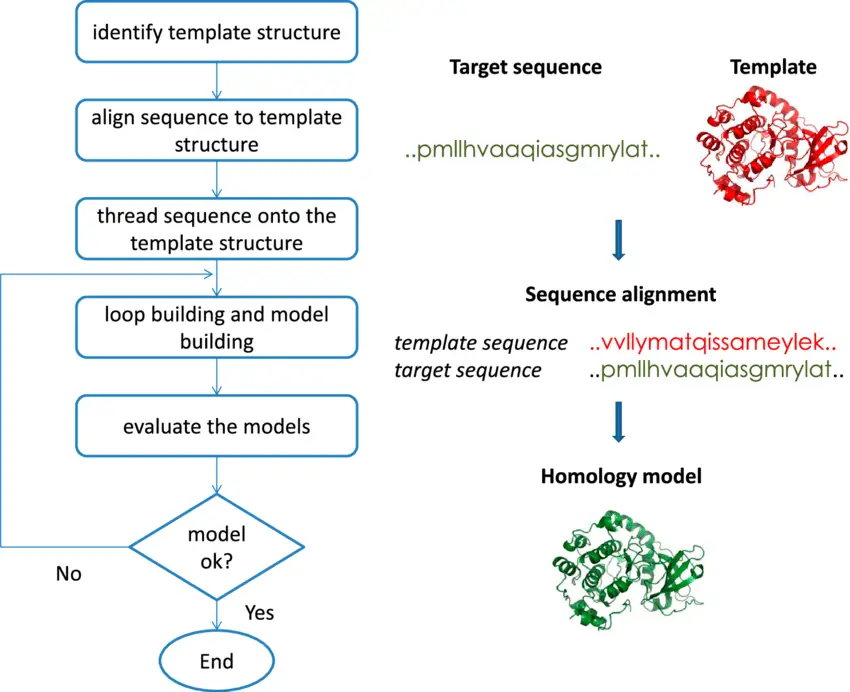
- Target Selection: The process of target selection involves the identification of a specific protein or macromolecule for which the 3D structure is to be predicted. The protein in question is commonly denoted as the target protein.
- Template Identification: Template Identification involves the identification of one or more template proteins that possess known experimental structures and exhibit a substantial sequence similarity with the target protein. The selection of templates must be conducted with caution to guarantee their structural pertinence to the protein of interest.
- Sequence Alignment: Conducting a sequence alignment is necessary to identify the corresponding amino acid positions between the target protein and the template protein(s). The process of alignment plays a crucial role in determining the association between the template and target sequences.
- Template Structure Preparation: Template Structure Preparation involves the preparation of template structures that will be utilized in the modeling process. The process may entail the elimination of extraneous constituents from the template architecture, such as water molecules or ligands, and verifying that the template structure conforms to the requisite format for modeling.
- Model Building: The process of model building involves the creation of an initial 3D model of a target protein through the transfer of structural information from template protein(s) to the target protein. This transfer is based on the alignment of sequence positions. The aforementioned procedure entails the adjustment of the template structure(s) to conform with the target sequence, followed by the placement of the corresponding residues in the model.
- Loop Modeling and Side-Chain Placement: The process of Loop Modeling and Side-Chain Placement involves the modeling of any absent loops or flexible regions within the protein structure being studied. This stage entails the anticipation of the conformation of loop regions that are absent in the template configuration. Furthermore, it is recommended to position the side chains of the amino acids within the model in a manner that maximizes the local interactions and energetics.
- Model Refinement: Model refinement involves the process of enhancing the quality and accuracy of the initial model. The optimization of the model’s geometry and enhancement of its stability and overall quality can be achieved through various techniques such as energy minimization, molecular dynamics simulations, and other optimization methods.
- Model Evaluation and Validation: The process of Model Evaluation and Validation involves the assessment of the quality and reliability of the final model. Validation of the model can be accomplished through diverse methods and criteria, such as assessing the model’s energy, verifying proper stereochemistry, and cross-referencing the model against available experimental data. The process of validation serves the purpose of verifying the biological plausibility and reliability of the model for subsequent analysis.
Advantages of Homology Modeling
- Prediction of Protein Structures: Homology modeling permits the prediction of protein structures, yielding valuable insights into the three-dimensional organization of proteins. Even in the absence of experimental structural data, it aids in comprehending their folding, stability, and functional properties.
- Speed and Cost-Effectiveness: In comparison to experimental techniques such as X-ray crystallography or NMR spectroscopy, homology modeling is relatively quick and economical. It can provide structural information about a target protein in a reasonable amount of time, making it a useful instrument for researchers with limited resources.
- Insight into Protein Function: Homology modeling provides insight into protein function by comparing the structure of a target protein to that of a template protein whose function is known. It can facilitate the comprehension of protein-ligand interactions, protein-protein interactions, and the molecular mechanisms underlying particular biological processes.
- Accessibility: The software and tools for homology modeling are broadly accessible and available to researchers. Numerous of these tools have intuitive interfaces, making them accessible to users with limited computational modeling experience.
- Protein Engineering and Drug Design: Homology modeling serves an important role in protein engineering and drug design. It facilitates rational drug design by providing accurate models of target proteins for virtual screening and molecular docking studies, thereby aiding in the design and optimization of enzymes.
Limitations of Homology Modeling
- Sequence Similarity Requirement: Homology modeling requires the availability of a suitable template protein with a high degree of sequence similarity to the target protein. Homology modeling becomes difficult or unreliable if the target protein shares little or no sequence similarity with any known protein.
- Template Quality and Structural Variability: The precision of homology models relies significantly on the relevance and quality of the selected template structure. If the template protein’s structural information is of low quality or inaccurate, it may have a negative effect on the quality of the modeled structure. In addition, if the target protein has substantial structural differences from the template, homology modeling may not capture these differences accurately.
- Errors in Alignment and Model Building: Errors in Sequence Alignment and Model Construction: Errors in the sequence alignment between the target and template proteins can result in inaccurate modeling and misinterpretation of protein structures. Inaccuracies during model construction or refinement can also impact the quality of the final product.
- Loop Modeling and Insertions/Deletions: Homology modeling frequently struggles to accurately predict the structures of highly flexible and variable protein loop regions. Likewise, modeling insertions or deletions in the target protein sequence can be difficult and may introduce uncertainty into the modeled structure.
- Model Evaluation and Validation: It is essential to evaluate the quality and dependability of homology models. However, it can be difficult to accurately validate the models, particularly in the absence of readily available experimental validation methods. The accuracy of predicted structures may be affected by the use of computational assessment tools.
Applications of Homology Modeling
The technique of homology modeling, also known as comparative modeling, is utilized in diverse fields of biological investigation. Several significant applications of homology modeling include:
- Protein Structure Prediction: The technique of homology modeling is widely employed in the prediction of protein structure, particularly in cases where experimental approaches such as X-ray crystallography or NMR spectroscopy are impractical or require significant time investment. The analysis of the spatial configuration of atoms within a protein offers significant insights that facilitate comprehension of its folding mechanism, stability, and functional attributes.
- Drug Discovery and Design: Homology modeling is a pivotal aspect of drug discovery and design as it facilitates the generation of precise models of protein targets, including enzymes and receptors, which are utilized for virtual screening and rational drug design. This process facilitates the identification of prospective binding sites, comprehension of protein-ligand interactions, and enhancement of drug candidates to augment their potency and selectivity.
- Enzyme Engineering: Homology modeling is a widely employed technique in the realm of enzyme engineering, which enables the design and customization of enzymes to cater to specific applications. Through the anticipation of an enzyme’s configuration and the residues present in its active site, scholars can gain a deeper comprehension of its catalytic process and make knowledgeable adjustments to improve its efficacy, endurance, substrate specificity, or selectivity.
- Protein Function Prediction: The prediction of protein function can be facilitated through homology modeling, which involves the comparison of the sequence and structure of a protein with those of established proteins in order to gain valuable insights. In the event that a protein’s structure with a confirmed function is accessible, it can be utilized as a framework to anticipate the function of the target protein by drawing comparisons between their structures and active sites.
- Understanding Protein-Protein Interactions: The prediction of interfaces and binding modes of protein-protein interactions is accomplished through the utilization of homology modeling. Through the utilization of protein structure modeling and complex analysis, researchers can acquire valuable knowledge regarding the molecular foundation of protein interactions. This knowledge is essential for comprehending cellular signaling pathways and developing therapeutic interventions that specifically target protein-protein interactions.
- Evolutionary Studies: The utilization of homology modeling can be advantageous in the examination of the evolutionary connections among proteins within the field of evolutionary studies. Through the comparative analysis of protein structures and sequences across various species, scholars can deduce their shared lineage, detect conserved domains, and explore the functional implications of sequence divergences.
FAQ
What is homology modeling?
Homology modeling, also known as comparative modeling, is a computational technique used to predict the 3D structure of a protein or other macromolecule based on its amino acid sequence and the known structure of a related protein.
How does homology modeling work?
Homology modeling works by aligning the amino acid sequence of the target protein with a template protein of known structure. The aligned sequences are used to transfer structural information from the template to the target, resulting in a predicted 3D model of the target protein.
What are the prerequisites for successful homology modeling?
The success of homology modeling depends on the availability of a suitable template with a high sequence similarity to the target protein. Adequate sequence alignment and careful model refinement are also crucial for generating accurate models.
What are the limitations of homology modeling?
Homology modeling has limitations when there is low sequence similarity between the target and template proteins, or when the target protein has unique structural features not present in any known template. Errors in alignment and model building, and the challenge of accurately predicting loop regions, can also affect the quality of the models.
How accurate are homology models?
The accuracy of homology models varies depending on factors such as sequence similarity, template quality, and refinement techniques employed. In general, models with high sequence similarity to the template and proper refinement have good overall accuracy, especially in regions of high conservation.
Can homology modeling predict protein function?
Homology modeling can provide insights into protein function by comparing the structure of the target protein with that of a template protein of known function. The assumption is that proteins with similar structures often perform similar functions.
Can homology modeling be used for membrane proteins?
Homology modeling can be used for membrane proteins, although it can be more challenging compared to soluble proteins due to the unique features of membrane protein structures. Special considerations, such as incorporating information on transmembrane regions and lipid interactions, are necessary for accurate modeling.
What software or tools are commonly used for homology modeling?
Popular software tools for homology modeling include MODELLER, SWISS-MODEL, Rosetta, I-TASSER, and Phyre2. These tools provide user-friendly interfaces and various functionalities for the modeling process.
How can the accuracy of homology models be assessed?
The accuracy of homology models can be assessed through various methods, including assessing stereochemistry, evaluating energy profiles, comparing the models to experimental data (if available), and using quality assessment scores such as Ramachandran plots or MolProbity scores.
What are some applications of homology modeling?
Homology modeling has diverse applications, including protein structure prediction, drug discovery, enzyme engineering, protein-protein interaction studies, and evolutionary studies. It serves as a valuable tool for understanding protein structure-function relationships and guiding experimental investigations.


