Table of Contents
What is Plate Count Agar (PCA)?
- Plate Count Agar (PCA) is a type of bacteriological substrate used for quantifying the total number of viable, aerobic bacteria present in a sample. Unlike selective media that favor the growth of specific types of bacteria, PCA is a non-selective medium. The bacterial count is typically expressed as colony-forming units per gram (CFU/g) for solid samples or per milliliter (CFU/ml) for liquid samples.
- The recommended technique for using PCA is the pour plate technique. In this method, the sample is diluted, and appropriate dilutions are added to sterile Petri plates. Molten agar, which is sterile and liquefied, is then poured onto the plates containing the diluted sample. The plates are gently rotated to ensure thorough mixing of the sample with the agar. Subsequently, the plates are incubated at a temperature of either 20 or 30 degrees Celsius for a duration of three days.
- After the incubation period, the colonies that have developed on the plates are counted. It is recommended to count colonies on plates that have 25 to 250 visible colonies, as this range is considered to provide the most accurate result. When determining the actual number of bacteria in the original sample, the dilution factor used during the pour plate technique must be taken into account.
- By using PCA and the appropriate technique, the total number of viable aerobic bacteria present in a sample can be determined, allowing for assessment and monitoring of bacterial populations in various environments, including food, water, and clinical samples.
Principle of Plate Count Agar (PCA)
The principle of Plate Count Agar (PCA), also known as Tryptone Glucose Yeast Agar or Casein-Peptone Dextrose Yeast Agar, is based on providing a nutrient-rich medium to support the growth of aerobic bacteria and facilitate their enumeration.
The PCA medium consists of various components that contribute to bacterial growth. Enzymatic digest of casein serves as a source of amino acids, nitrogen, carbon, vitamins, and minerals, providing essential nutrients required for bacterial metabolism and multiplication. Yeast extract is another key ingredient that supplies B-complex vitamins, which are important cofactors for many enzymatic reactions involved in bacterial growth.
Glucose, a fermentable carbohydrate present in PCA, acts as an energy source for bacterial metabolism. Bacteria can utilize glucose through fermentation, producing energy and metabolic byproducts. The availability of glucose ensures that bacteria have the necessary nutrients for growth and reproduction.
Agar, a solidifying agent derived from seaweed, is added to the PCA medium. Agar forms a gel-like matrix when cooled, providing a solid surface for bacterial colonies to develop. It helps in the isolation and enumeration of individual colonies, facilitating the quantification of bacterial populations.
Overall, the principle of PCA relies on providing a nutrient-rich environment containing amino acids, vitamins, minerals, and fermentable carbohydrates to support the growth of aerobic bacteria. The solidifying agent, agar, allows for the visualization and enumeration of individual colonies formed by bacteria, enabling the determination of viable bacterial counts in a sample.
Composition of Plate Count Agar (PCA)
| Ingredients | Gms/L |
| Enzymatic Digest of Casein/tryptone | 5.0 |
| Yeast Extract | 2.5 |
| Glucose | 1.0 |
| Agar | 15.0 |
Final pH 7.0 ± 0.2 at 25°C
Preparation of Plate Count Agar (PCA)
The preparation of Plate Count Agar (PCA) involves the following steps:
- Weigh and Suspend: Measure 23.5 grams of Plate Count Agar powder and suspend it in 1000 ml of distilled water. The water should be at room temperature.
- Heat and Dissolve: Heat the suspension to boiling while stirring continuously to ensure complete dissolution of the medium. Continue boiling until all the components are dissolved.
- Autoclave Sterilization: Sterilize the PCA medium by autoclaving. Place the medium in an autoclave and subject it to a pressure of 15 pounds per square inch (psi) or 121°C for 15 minutes. Autoclaving helps eliminate any potential contaminants and ensures a sterile medium.
- Cooling: After autoclaving, allow the PCA medium to cool down to a temperature of 45-50°C. This temperature range ensures that the medium is still in a liquid state and can be easily poured into Petri plates.
- Mixing and Pouring: Once the PCA medium has reached the desired temperature, mix it well to ensure homogeneity. Gently swirl the container to achieve uniformity. Then, pour the medium into sterile Petri plates, making sure to cover the bottom of the plates evenly.
It is important to maintain aseptic conditions throughout the preparation process to avoid contamination. After pouring the PCA medium into Petri plates, allow the plates to solidify before use. Once solidified, the PCA plates are ready to be used for bacterial enumeration and other microbiological analyses.
Spread plate Method
The spread plate method is a widely used technique in microbiology for the enumeration and isolation of bacteria from a sample. Here are the steps involved in the spread plate method:
- Drying the Petri Dish: Before starting the procedure, the Petri dishes should be dried in an oven with the lids slightly ajar. This helps remove any moisture that could affect the spreading of the inoculum.
- Sample Dilution: Prepare a series of decimal dilutions of the sample to be analyzed. Starting with the original sample, transfer 0.1 mL of the sample to the first dilution tube, and then transfer 0.1 mL from the first dilution tube to the second, and so on. This creates a dilution series that allows for the enumeration of bacterial colonies within the countable range.
- Inoculation: Using a sterile spreader, transfer a small volume (usually 0.1 mL) of the sample from each dilution onto the surface of a solid agar medium contained in a Petri dish. The spreader is gently moved back and forth across the surface of the agar, evenly distributing the inoculum.
- Incubation: After inoculating the samples, the Petri dishes are incubated in a controlled environment at a temperature of 30 ± 1°C for 72 ± 3 hours. This specific incubation period allows for optimal bacterial growth and colony formation.
During incubation, the bacteria present in the sample will grow and form visible colonies on the agar surface. The colonies will be distributed across the plate, with each colony representing a single viable bacterium present in the original sample.
After incubation, the plates are examined, and bacterial colonies are counted. The results are reported as colony-forming units (CFUs) per milliliter or gram, taking into account the dilution factors used during the procedure.
The spread plate method is widely used for bacterial enumeration, isolation of pure cultures, and assessment of bacterial growth characteristics. It provides a quantitative measure of viable bacteria in a sample and allows for the isolation of individual colonies for further characterization and identification.
Test Results on Plate Count Agar (PCA)
When analyzing results on Plate Count Agar (PCA), the following steps can be followed:
- Colony Count: Count the number of visible colonies on the PCA plates. It is recommended to count colonies on plates that have a colony count ranging from 15 to 300. This range ensures optimal accuracy in the enumeration of bacterial colonies.
- Calculation of CFU/ml or CFU/g: The count of colonies is reported as Colony-Forming Units (CFU) per milliliter (CFU/ml) for liquid samples or per gram (CFU/g) for solid samples. To calculate the actual CFU/ml or CFU/g, the dilution factor used during the sample preparation and plating must be taken into consideration. Multiply the colony count by the appropriate dilution factor to obtain the final CFU/ml or CFU/g result.
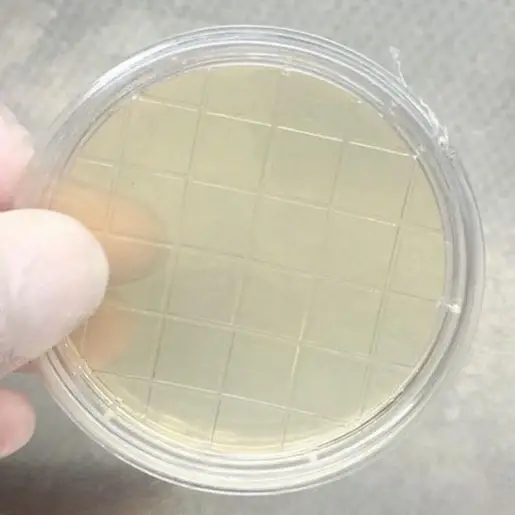
- Identification of Colonies: Based on the growth observed on the PCA plates, identify the types of colonies present. In this example, the growth of Bacillus subtilis, Escherichia coli, and Staphylococcus aureus is mentioned, with all colonies appearing as straw-colored. Different bacterial species may exhibit characteristic colony morphologies and colors, aiding in identification.
- Recording and Reporting: Record the colony counts and the identification of the bacterial species observed on the PCA plates. Report the results in CFU/ml or CFU/g, along with any relevant dilution factors used during sample preparation.
| Bacteria | Growth |
|---|---|
| Staphylococcus aureus | Good growth, productivity PR ≥ 0.70 |
| Escherichia coli | Good growth, productivity PR ≥ 0.70 |
| Bacillus subtilis | Good growth, productivity PR ≥ 0.70 |
By following these steps, the results obtained from PCA plates provide an estimation of the total number of viable aerobic bacteria in the original sample. These results are crucial for various applications, such as assessing the microbial quality of food, water, and clinical specimens, as well as monitoring the effectiveness of antimicrobial treatments.
Quality Control
Quality control measures are essential in microbiology to ensure the consistency and reliability of laboratory procedures and results. Here are some aspects of quality control related to the characteristics and cultural response of the medium:
- Appearance: The quality control assessment includes checking the appearance of the medium. In this case, the Plate Count Agar (PCA) appears as a cream to yellow homogeneous free-flowing powder.
- Gelling: The gelling property of the medium is evaluated. It is expected to form a firm gel comparable to a 1.5% Agar gel. This characteristic ensures that the medium solidifies adequately to support bacterial growth and colony formation.
- Colour and Clarity: The quality control assessment involves observing the prepared medium in Petri plates. It should form a light yellow-colored clear to slightly opalescent gel. This characteristic indicates the proper preparation and clarity of the medium.
- Reaction and pH: The reaction of a 2.35% w/v aqueous solution of the medium is evaluated. The pH of the solution should be within the range of 6.80-7.20 at 25°C. This pH range ensures optimal conditions for bacterial growth and metabolic activity.
- Organism Inoculum and Cultural Response: To assess the medium’s performance, specific bacterial strains are inoculated onto the medium and evaluated for growth and recovery. The cultural response is determined by incubating the inoculated medium at 35-37°C for 18-48 hours.
The provided data shows the expected growth and recovery of various bacterial strains, such as Enterococcus faecalis, Escherichia coli, Lactobacillus casei, Staphylococcus aureus subsp. aureus, and Streptococcus pyogenes. Luxuriant growth, defined as a significant number of visible colonies, is expected, with a minimum recovery rate of 70% for each strain.
Uses of Plate Count Agar (PCA)
Plate Count Agar (PCA) has several uses in the field of microbiology. Here are some common applications:
- Enumeration of Bacteria: PCA is widely used for the quantitative enumeration of bacteria in various samples, including food, water, and other materials of sanitary importance. By plating the samples on PCA, viable aerobic bacteria can form visible colonies, allowing for their counting and estimation of bacterial population density.
- Microbial Quality Assessment: PCA helps assess the microbial quality of food products. It is employed to determine the total viable bacterial count, which serves as an indicator of hygiene and potential spoilage. Monitoring bacterial levels in food samples can aid in quality control, shelf-life determination, and identification of potential microbial contaminants.
- Water Quality Analysis: PCA is also used in water quality analysis to determine the level of bacterial contamination. By analyzing water samples using PCA, the total bacterial count can be obtained, providing valuable information about water safety and potential risks to human health.
- Environmental Monitoring: PCA can be utilized to enumerate bacteria in sterile rooms or controlled environments. This helps assess the microbial cleanliness of surfaces and air, aiding in monitoring and maintaining sterile conditions in healthcare facilities, pharmaceutical industries, and research laboratories.
- Research and Educational Purposes: PCA is commonly used in research and educational settings for microbiological studies. It provides a simple and reliable method for quantifying bacterial populations, allowing researchers and students to investigate microbial growth patterns, perform antimicrobial efficacy studies, and evaluate the impact of various environmental factors on bacterial growth.
Limitations of Plate Count Agar (PCA)
While Plate Count Agar (PCA) is a widely used medium for bacterial enumeration, it does have certain limitations. Here are some limitations of PCA:
- Limited Identification: PCA is primarily used for quantifying bacterial populations based on colony counts. However, it does not provide specific identification of individual bacterial species. For complete identification and characterization of bacterial isolates, further testing using biochemical, immunological, molecular, or mass spectrometry methods is recommended.
- Non-Selective Nature: PCA is a non-selective medium, meaning it supports the growth of a wide range of aerobic bacteria. While this broad applicability is advantageous for general bacterial enumeration, it may not be suitable for the growth of fastidious organisms that have specific nutritional requirements or growth preferences. Some bacteria with complex nutritional needs or slow growth rates may not thrive well on PCA.
- Bias Towards Aerobic Bacteria: PCA is specifically designed to support the growth of aerobic bacteria, which may lead to an underestimation of anaerobic bacterial populations present in a sample. Anaerobic bacteria require specialized conditions and media for growth, and their enumeration may require the use of specific anaerobic culture methods.
- Limited Differential Characteristics: PCA does not contain specific indicators or components that allow for the differentiation of bacterial species based on specific biochemical or physiological characteristics. It primarily provides a supportive growth environment, allowing bacterial colonies to form without providing distinct visual or biochemical markers for species differentiation.
- Lack of Selectivity: PCA does not contain inhibitors or selective agents that can suppress the growth of specific bacteria or favor the growth of certain types of bacteria. This may limit its usefulness for specific applications where the focus is on a particular group of bacteria or the inhibition of unwanted bacterial contaminants.
FAQ
What is Plate Count Agar (PCA)?
Plate Count Agar (PCA) is a bacteriological substrate used for the enumeration of viable aerobic bacteria in a sample.
What is the purpose of using PCA?
PCA is primarily used to determine the total number of live aerobic bacteria present in a sample. It provides a suitable growth medium for bacterial colonies to form and be counted.
Is PCA selective or differential?
PCA is a non-selective medium, meaning it supports the growth of a wide range of aerobic bacteria. It is not a differential medium as it does not contain specific indicators for distinguishing different bacterial species.
How is PCA prepared?
PCA is typically prepared by dissolving the PCA powder in distilled water, followed by sterilization through autoclaving. The sterile medium is then poured into Petri dishes and allowed to solidify before use.
How are samples plated on PCA?
To plate samples on PCA, a known volume of the sample is spread evenly over the surface of the solidified PCA medium using a sterile spreader. This allows the bacteria to grow and form visible colonies.
What is the incubation period for PCA?
PCA plates are typically incubated at a controlled temperature, usually around 30 ± 1°C, for a specified period, such as 72 ± 3 hours. This allows sufficient time for bacterial colonies to develop and become visible.
How are colonies counted on PCA plates?
After incubation, the plates are examined, and visible bacterial colonies are counted. It is recommended to count plates with colony numbers ranging from 15 to 300, as this range provides the most accurate results.
Can PCA be used for anaerobic bacteria?
PCA is primarily designed for the enumeration of aerobic bacteria and may not support the growth of anaerobic bacteria effectively. Specialized media and conditions are required for the enumeration of anaerobic bacteria.
Can PCA identify specific bacterial species?
PCA alone does not provide specific identification of individual bacterial species. Further tests, such as biochemical, molecular, or immunological techniques, are needed for precise identification.
What are the limitations of PCA?
Limitations of PCA include its non-selective nature, which may not support the growth of fastidious organisms, and the lack of differentiation between different bacterial species. Additionally, it does not provide information on the viability or metabolic activity of bacteria.
References
- FDA – Aerobic Plate Count
- Standard Methods for the Examination of Dairy Products
- Handbook of Culture Media for Food Microbiology
- FDA – Plate Count Agar (Standard Methods)
- Pierre-Yves Guillaume – GELOSE POUR DENOMBREMENT PCA
- Indicia fiche technique : Gélose PCA
- Biokar – GELOSE POUR DENOMBREMENT (PCA
- neogen – Plate Count Agar (Standard Methods)
- https://www.clinisciences.com/en/buy/cat-plate-count-agar-pca-non-selective-5497.html
- http://www.vetbact.org/displayextinfo/97
- https://www.neogen.com/categories/microbiology/plate-count-agar-standard-methods/
- https://www.thermofisher.com/order/catalog/product/R09642
- https://www.grosseron.com/oo/Assets/client/FTP/GROSSERON/FT/FT9010032.pdf
- https://www.humeau.com/media/blfa_files/__Fiche_de_spe__cification_plate-count-agar-4×5-btes-90mm-18010077402_EN_ea30d0f96cc23c0fc065c4200ef7e9ca_217317.pdf
- https://exodocientifica.com.br/_technical-data/M091.pdf
- https://microbiologie-clinique.com/pca-agar-plate-count.html
