Table of Contents
In 1953, James Watson and Francis Crick made a groundbreaking contribution to the understanding of DNA structure. By combining physical and chemical data, they proposed a model for DNA as a double helix, a twisted molecule consisting of two complementary strands held together by hydrogen bonds. This model revolutionized our knowledge of genetics and unlocked the genetic code that underlies all living organisms.
The Watson and Crick model revealed that DNA serves as the backbone for all life forms. It demonstrated that the structure of DNA contains genetic information and can make copies of itself, which are then passed down through generations. This model provided a key insight into how DNA functions and plays a fundamental role in the transmission of genetic traits.
The Watson and Crick model, also known as the double helix model, represented a scientific breakthrough of immense significance. It explained that DNA is composed of two strands, each consisting of a sugar-phosphate backbone and nitrogenous bases. These bases, including adenine, thymine, guanine, and cytosine, pair up through hydrogen bonds to form the rungs of the DNA ladder. This model not only explained the structure of DNA but also how it replicates and carries out essential biological processes.
Today, the Watson and Crick model remains a fundamental concept in molecular biology. It provides one of the best visual representations of the structure of double-helix DNA. DNA itself is a polymer composed of monomer units called deoxyribonucleotides, which are linked together by phosphodiester bonds. Many scientists, including Friedrich Miescher, P.A. Levene, W.T. Astbury, Maurice Wilkins, and Rosalind Franklin, contributed to the understanding of DNA’s components and composition, laying the groundwork for Watson and Crick’s breakthrough.
The significance of Watson and Crick’s model was widely recognized, leading to their receipt of the Nobel Prize in Physiology or Medicine in 1962. They shared this prestigious award with Maurice Wilkins, although Rosalind Franklin, who made significant contributions to the understanding of DNA structure, tragically passed away in 1958 and thus did not receive the recognition she deserved.
History of DNA Double Helical Structure
The history of the DNA double helical structure is a remarkable journey of scientific discovery that involved several key scientists and breakthroughs. Here is a timeline highlighting the significant events:
1869: Freidrich Miescher, a Swiss physiological chemist, identified a substance in the nuclei of white blood cells, which he named “Nuclein.” This discovery laid the foundation for the understanding of nucleic acids, such as deoxyribonucleic acid (DNA).
1920: Phoebus Aaron Theodore Levene, an American organic chemist, determined the elemental composition of nucleic acids. He proposed the “Tetranucleotide hypothesis,” suggesting that the base composition of all four nitrogenous bases (A, T, G, and C) would be the same.
1940: William Thomas Astbury, an English physicist and molecular biologist, utilized X-ray crystallography to present a three-dimensional model of DNA. This model provided insights into the structural characteristics of DNA.
1950: Erwin Chargaff, an Austro-Hungarian biochemist, made significant contributions to the understanding of DNA biochemistry. He formulated two key principles known as “Chargaff’s Rule.” The first principle stated that the sum of purines (A and G) is equal to the sum of pyrimidines (T and C) in DNA. The second principle highlighted variations in DNA composition between different species.
1952: Maurice Wilkins, Rosalind Franklin, and their colleagues utilized X-ray diffraction to obtain photographs of DNA. Rosalind Franklin’s super X-ray diffraction photograph provided crucial insights into the structure of DNA.
1953: James Watson and Francis Crick, working in collaboration with Wilkins and Franklin, unveiled the double-helical structure of DNA. Watson and Crick’s model, based on the X-ray diffraction data, provided a groundbreaking understanding of how DNA is structured. This discovery was widely acclaimed and earned them the Nobel Prize in Physiology or Medicine in 1962, which they shared with Wilkins.
The discovery of the DNA double helical structure revolutionized our understanding of genetics and paved the way for advancements in molecular biology. The contributions of these scientists laid the foundation for further research in genetics, DNA replication, and the decoding of the genetic code, making significant contributions to the field of biological sciences.
4 Key features of Watson and Cricks DNA model
The Watson and Crick model of DNA, which revolutionized our understanding of genetics, is characterized by four major features:
- Complementary Base Pairing: The DNA molecule consists of two individual strands of polynucleotides that are connected by hydrogen bonds. The nucleotide bases in DNA, adenine (A), thymine (T), guanine (G), and cytosine (C), exhibit a complementary pairing rule: purine bases (A and G) always pair with pyrimidine bases (T and C). This base pairing allows for the precise replication and transmission of genetic information.
- Right-Handed Double Helix: In most cases, the double helix structure of DNA is right-handed, meaning it twists in a clockwise direction when viewed along its axis. It completes one full turn every 34 angstroms (34A°). Visualizing the double helix, if you hold your right thumb pointed up to represent the axis, the sugar-phosphate backbones run along the outside of the molecule, while the nitrogen bases are positioned inside. An example of a left-handed DNA structure is the Z-DNA, which is an exception to the right-handed helix.
- Anti-Parallel Strands: The two strands of DNA in the double helix are anti-parallel to each other. This means that if one strand has a starting point at the 5′ end, the other strand will have its starting point at the 3′ end, and vice versa. This arrangement is due to the polarity of the phosphodiester linkage, which connects the nucleotides. The anti-parallel nature of the strands is crucial for DNA replication and transcription processes.
- Hydrogen Bonding and Accessibility: The outer exposed molecules of the nitrogen bases in DNA have the potential to form hydrogen bonds with other molecules. These hydrogen bonds play a crucial role in DNA replication and gene expression. They provide easy access to the DNA molecule, allowing enzymes and proteins to interact with specific base sequences for processes such as DNA replication, transcription, and protein synthesis.
These four major features of the Watson and Crick DNA model—complementary base pairing, right-handed double helix, anti-parallel strands, and hydrogen bonding—provide a comprehensive understanding of the structure and function of DNA. They explain how genetic information is stored, replicated, and expressed, and have paved the way for significant advancements in the field of molecular biology.
Components of DNA
DNA, or deoxyribonucleic acid, is composed of specific components that together form the structure of the molecule. These components include nucleotide monomers, which are the building blocks of DNA.
- The nucleotide monomers in DNA consist of three molecular parts: nitrogen bases, deoxyribose sugar, and a phosphate group. There are four types of nitrogen bases found in DNA: adenine (A), cytosine (C), thymine (T), and guanine (G). These bases are responsible for carrying the genetic information in the DNA molecule.
- The deoxyribose sugar molecule is a five-carbon sugar that forms the backbone of the DNA strand. Each nucleotide in the DNA chain is connected to the adjacent nucleotide through the deoxyribose sugar. The deoxyribose sugar provides structural stability to the DNA molecule.
- The phosphate group is attached to the deoxyribose sugar in each nucleotide. It consists of a phosphorus atom bonded to four oxygen atoms. The phosphate groups link together to form a phosphate backbone, running along the outer edges of the DNA strands. The phosphate backbone provides additional structural support and stability to the DNA molecule.
In summary, the components of DNA include nucleotide monomers, which consist of nitrogen bases (adenine, cytosine, thymine, and guanine), deoxyribose sugar, and a phosphate group. These components combine to form the unique structure of DNA and encode the genetic information that is essential for all living organisms.
The Nitrogen Bases or Nucleotides
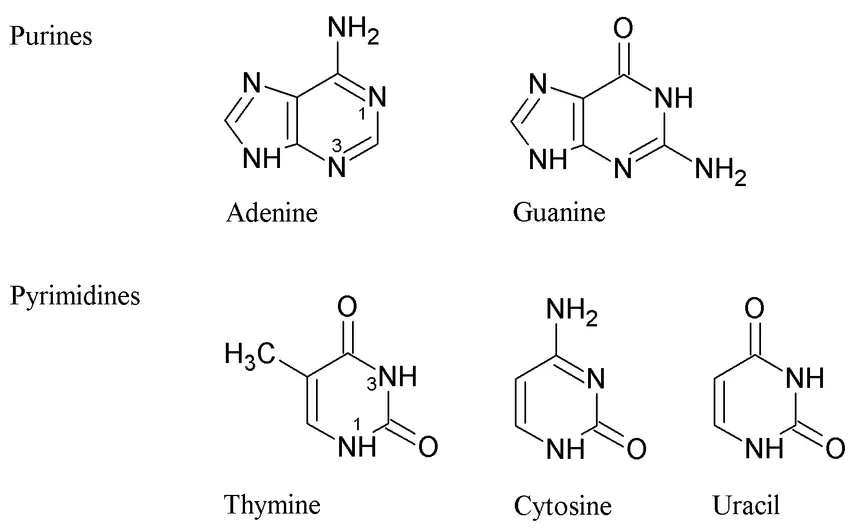
- The nitrogen bases, or nucleotides, play a crucial role in the structure and function of DNA. DNA strands are made up of these monomers, which are often referred to as bases due to their cyclic organic nature.
- There are four different nucleotides in DNA: adenine (A), thymine (T), cytosine (C), and guanine (G). These nucleotides join together to form a DNA strand, with the base portions projecting inward from the backbone of the strand.
- In the DNA double helix structure, two strands bind together via the nitrogen bases and twist around each other, forming a double helix. The nitrogen bases exhibit a specific pairing pattern.
- Adenine always pairs with thymine (A-T) via two hydrogen bonds, while cytosine always pairs with guanine (C-G) via three hydrogen bonds. This complementary base pairing ensures that the amount of adenine is equal to the amount of thymine, and the amount of guanine is equal to the amount of cytosine. The hydrogen bonding between the base pairs stabilizes the structure of the DNA double helix.
- The nitrogen bases in DNA are divided into two categories: purines and pyrimidines. Adenine and guanine are purines, while cytosine and thymine are pyrimidines. These bases form the foundation of the nucleotides and ultimately the DNA molecule.
- The shape of the DNA helix is stabilized by hydrogen bonding and hydrophobic interactions between the bases. The hydrogen atoms of the amino groups act as donors, while the carbonyl oxygen and ring nitrogen act as acceptors in the hydrogen bonding interactions.
- The DNA double helix structure contains ten nucleotides in each turn, with an internucleotide distance of 3.4 angstroms (3.4A°). The full turn of the helix has a length of 34 angstroms (34A°) and a diameter of approximately 20 angstroms (20A°).
- This helical structure gives rise to two grooves: the major groove and the minor groove. The major groove is deep and wide, serving as the specific binding site for proteins involved in DNA interactions. On the other hand, the minor groove is the narrower space between the two strands.
- In summary, the nitrogen bases or nucleotides in DNA are crucial components that contribute to the structure and function of the molecule. They form complementary base pairs, have specific hydrogen bonding patterns, and stabilize the DNA double helix.
- The nucleotide bases adenine, thymine, cytosine, and guanine are fundamental to the genetic code and play a vital role in DNA replication, transcription, and protein synthesis.
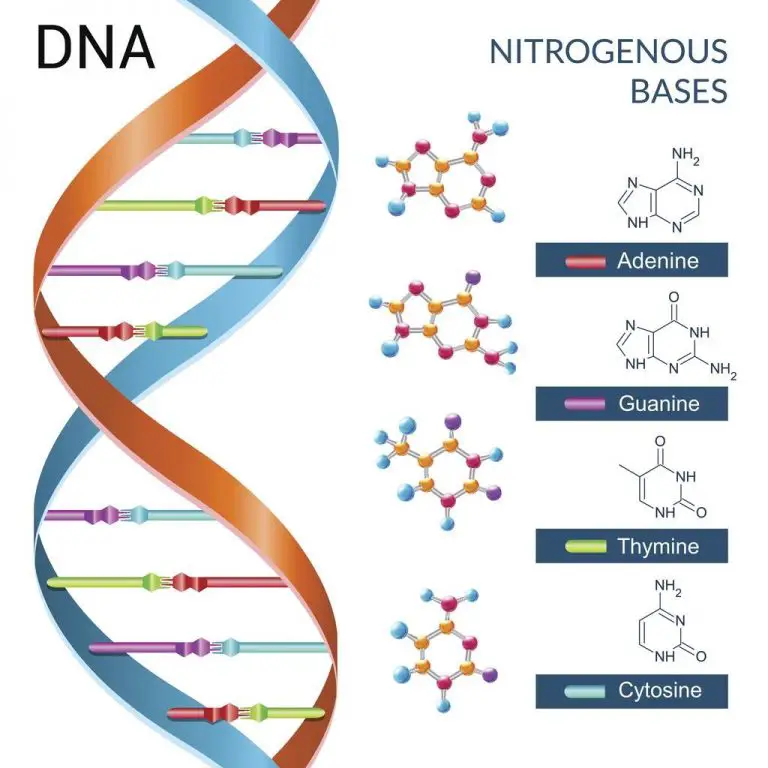
Deoxyribose Sugar
- Deoxyribose sugar, also known as 2-deoxyribose, is a pentose sugar that serves as a key component of deoxyribonucleic acid (DNA). It is derived from the pentose sugar ribose and has the chemical formula C5H10O4.
- In DNA, deoxyribose sugar forms the backbone of the molecule, alternating with phosphate groups. It binds to the nitrogenous bases adenine, thymine, guanine, and cytosine, which are responsible for carrying the genetic information in DNA. The absence of one hydroxyl group on the 2′ carbon atom distinguishes deoxyribose from ribose.
- Deoxyribose sugar plays a critical role in DNA, which represents the genetic information in all living cells. It is an essential component of life and is shared among all living organisms.
- The deoxyribose sugar molecule consists of five carbon atoms, with four of them forming carbon molecules and one oxygen molecule arranged cyclically. Its flexible structure allows it to adopt various conformations, such as the C2′ endo configuration observed in the canonical B-DNA.
- When a nucleobase is attached to a deoxyribose sugar, it forms a nucleoside. The 1′ carbon of the pentose sugar bonds with the nitrogenous base, while the 5′ carbon atom bonds to the phosphate group. These bonds are facilitated by hydrogen bonding.
- During DNA replication, enzymes are involved in breaking the hydrogen bonds, resulting in the separation of the DNA strands. The newly formed deoxyribose molecules then attach to nitrogenous bases and phosphate groups, ultimately forming a new DNA molecule. The process of replication ensures the accurate transmission of genetic information from one generation to the next.
- In summary, deoxyribose sugar is a vital component of DNA. It forms the backbone of the DNA molecule, binding to nitrogenous bases and phosphate groups. Its unique structure and bonding patterns contribute to the stability and functionality of DNA, enabling the storage and transmission of genetic information.
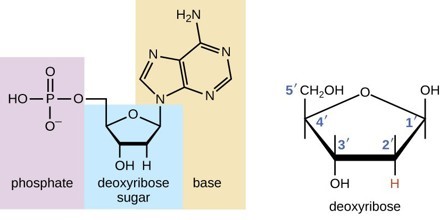
The Phosphate Group (Phosphate Backbone)
- The phosphate group, also known as the phosphate backbone, plays a critical role in the structure and function of nucleic acids, including DNA. It forms the structural framework of DNA, providing stability and defining the directionality of the molecule.
- The sugar-phosphate backbone is composed of alternating sugar and phosphate groups. In DNA, nucleotides are linked together in a chain by ester bonds between the sugar base of one nucleotide and the phosphate group of the adjacent nucleotide. The sugar is referred to as the 3′ end, while the phosphate is the 5′ end of each nucleotide.
- The phosphate group attached to the 5′ carbon of the sugar on one nucleotide forms an ester bond with the free hydroxyl group on the 3′ carbon of the next nucleotide. These bonds are known as phosphodiester bonds. As DNA is synthesized, the sugar-phosphate backbone extends or grows in the 5′ to 3′ direction.
- In double-stranded DNA, the sugar-phosphate backbones of the two strands run opposite each other, and they twist together to form the characteristic double-helix shape. This twisting and alignment of the sugar-phosphate backbones contribute to the stability and structural integrity of DNA.
- The phosphate group carries a negative charge, making the sugar-phosphate backbone negatively charged as well. This negative charge and the hydrophilic nature of the phosphate group allow the DNA backbone to interact with water molecules.
- The phosphate group is an essential component of every single strand of the DNA molecule. Its chemical structure consists of a phosphorus atom bonded with four oxygen atoms, with three single bonds and one double bond.
- Apart from its structural role, the phosphate group also functions as an energy donor during DNA synthesis. The release of phosphate groups provides energy for the formation of phosphodiester bonds between nucleotides.
- In summary, the phosphate group forms the backbone of DNA, linking nucleotides together through phosphodiester bonds. It provides stability, directionality, and the necessary negative charge for the molecule. The phosphate group plays a fundamental role in DNA structure, function, and the transmission of genetic information.
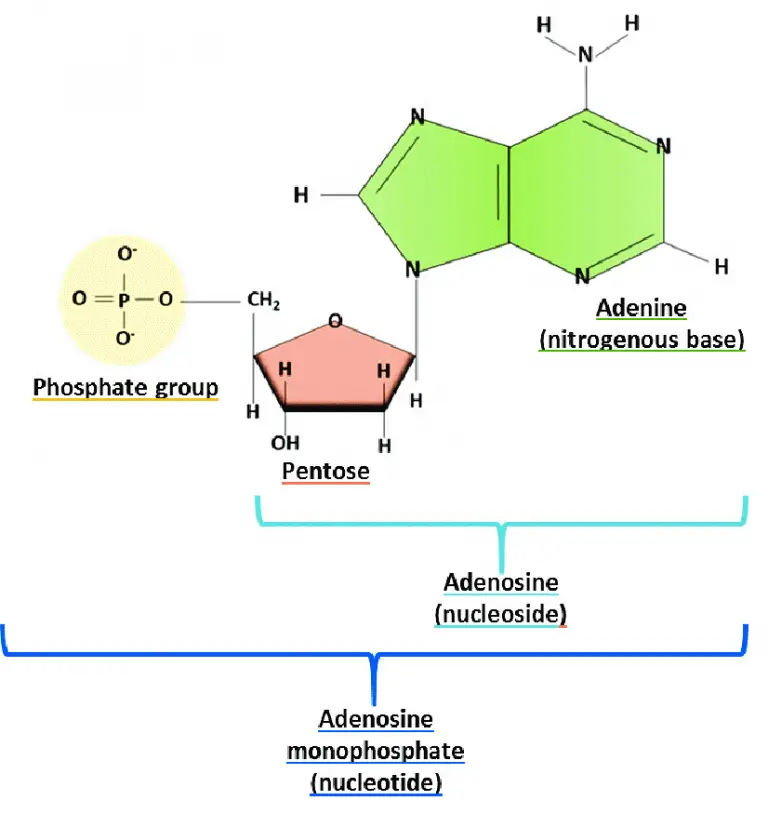
Watson and Crick model of DNA
Watson and Crick exhibited the structure of DNA after examining the manuscript of Linus Pauling and Corey. Linus Pauling and Corey presented the unsuccessful 3D structure of nucleic acid in 1953. Then, (in early 1953), Watson and Crick proposed a double-helical structure for DNA by combining physical and chemical property data. The main characteristics of the DNA model developed by Watson and Crick include:
Physical Properties of DNA
DNA, according to the Watson and Crick model, exhibits several physical properties that contribute to its structure and function:
- Double-stranded helix: DNA consists of two polynucleotide chains that are spirally twisted around each other, giving it a twisted ladder-like appearance. This double-stranded helical structure provides stability to the DNA molecule.
- Antiparallel polarity: The two polynucleotide strands of DNA run in opposite directions, resulting in antiparallel polarity. One strand runs in the 5′-3′ direction, while the other runs in the 3′-5′ direction. This arrangement allows for complementary base pairing between the strands.
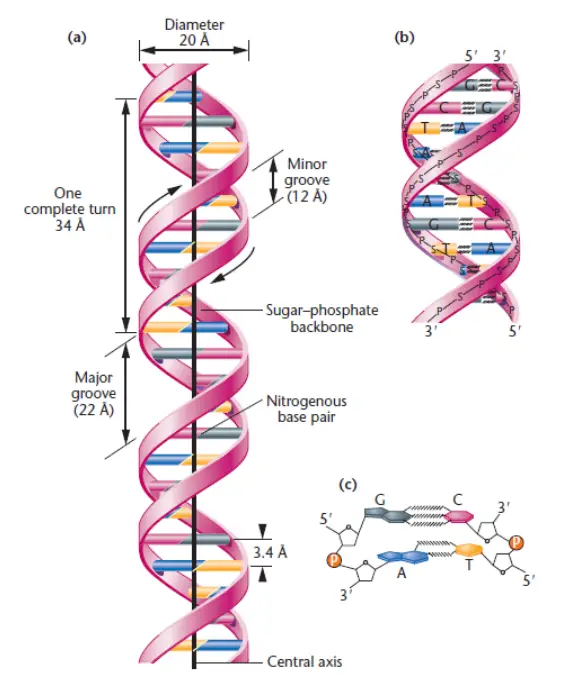
- Diameter: The diameter of the DNA double helix is approximately 20Å (angstroms). This consistent diameter is maintained due to the specific pairing of nucleotide bases within the helix.
- Internucleotide distance: The distance between two adjacent nucleotides, known as the internucleotide distance, is approximately 3.4Å. This distance refers to the spacing between the individual base pairs along the DNA molecule.
- Length of helix and base pairs per turn: A complete turn of the DNA helix has a length of approximately 34Å and consists of 10 base pairs. This characteristic length and base pairing arrangement contribute to the overall stability and structure of DNA.
- Right-handed twist: DNA exhibits a right-handed twist or a clockwise direction when viewed from the top. This twist allows the two strands to wrap around each other in a helical manner.
- Major and minor grooves: The twisting of the DNA strands creates distinct features known as major and minor grooves. The major groove is wider, while the minor groove is narrower. These grooves provide binding sites for various proteins involved in DNA replication, transcription, and other cellular processes.
Understanding the physical properties of DNA, such as its helical structure, diameter, and grooves, is crucial for comprehending its role in genetic information storage, replication, and gene expression. These properties enable DNA to function as the fundamental molecule of heredity in living organisms.
Chemical Properties of DNA
DNA exhibits various chemical properties that are crucial to its structure and function:
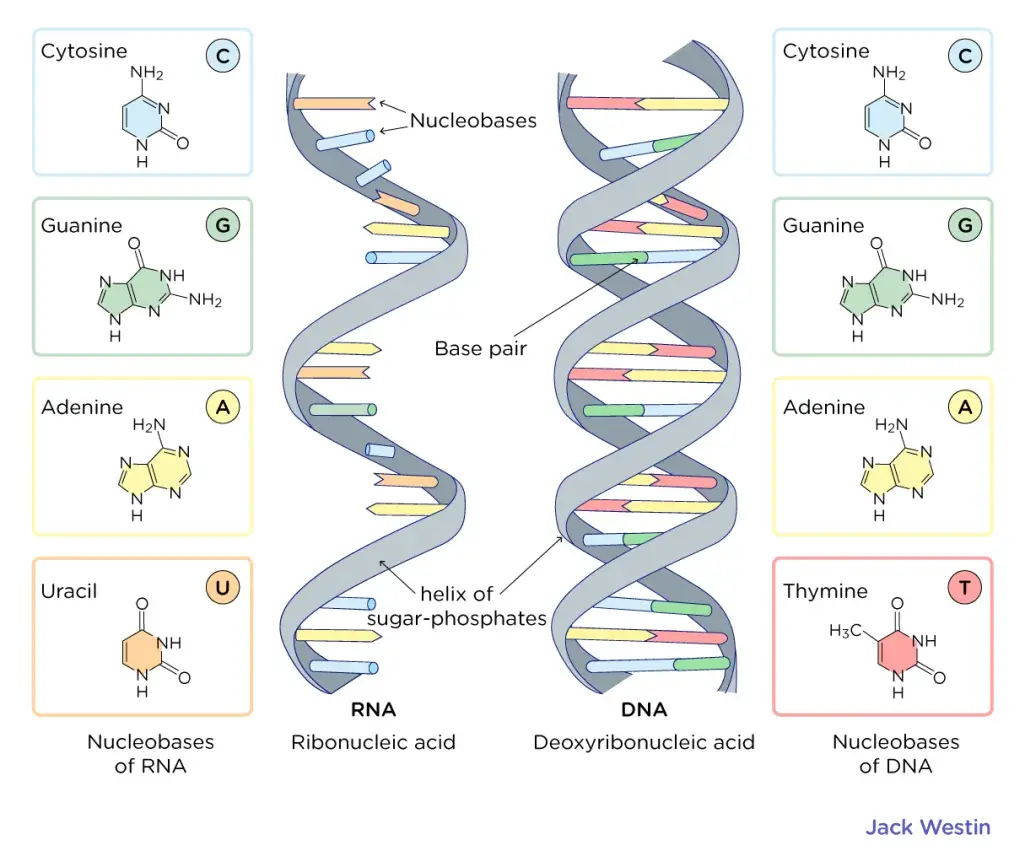
- Nucleotide bases: DNA is composed of four nucleotide bases: adenine (A), guanine (G), cytosine (C), and thymine (T). Adenine and guanine are purine bases, characterized by a single-ring structure, while cytosine and thymine are pyrimidine bases, featuring a double-ring structure.
- Complementary base pairing: The two DNA strands are joined together through complementary base pairing of the nucleotide bases. Adenine pairs with thymine through two hydrogen bonds, while guanine pairs with cytosine through three hydrogen bonds. This specific base pairing forms the foundation of DNA’s double-stranded structure.
- Hydrogen bonding: The nucleotide bases within the DNA strands are held together by strong hydrogen bonds. These hydrogen bonds contribute to the stability and integrity of the DNA molecule.
- Chargaff’s rule: The base composition of DNA follows Chargaff’s rule, which states that the sum of purines (A and G) is equal to the sum of pyrimidines (C and T). Thus, A + G = T + C. However, the individual base compositions of A + T and G + C may vary.
- Sugar-phosphate backbone: The backbone of DNA consists of a sugar-phosphate backbone, which forms the structural framework of the DNA molecule. The sugar component is deoxyribose, and it is linked to the phosphate group through phosphodiester bonds. The sugar-phosphate backbone provides stability and support to the DNA strands.
- DNA stability: The combination of phosphodiester bonds in the backbone and hydrogen bonds between the nucleotide bases contributes to the overall stability of DNA. These chemical interactions help maintain the double-stranded structure of DNA and protect the genetic information encoded within.
Understanding the chemical properties of DNA, such as complementary base pairing, hydrogen bonding, and the sugar-phosphate backbone, is essential for comprehending DNA replication, transcription, and the role of DNA in storing and transmitting genetic information. These chemical properties enable DNA to serve as the blueprint for life and play a vital role in cellular processes.
Features of the Watson-Crick Structure of DNA
The Watson-Crick model of DNA, which describes the structure of the DNA molecule, exhibits several distinct features:
- Double Helix: The DNA molecule adopts a right-handed double helix structure. It consists of two polynucleotide strands that coil around each other in a spiral fashion.
- Antiparallel Orientation: The two strands of DNA run in opposite directions, referred to as antiparallel. One strand has a 5′ end (with a phosphate group attached to the 5′ carbon of the sugar) and a 3′ end (with a hydroxyl group attached to the 3′ carbon), while the other strand has a reversed orientation.
- Exterior Sugar-Phosphate Backbone: The sugar-phosphate backbones of the DNA strands are positioned on the exterior of the helix. These backbones are composed of alternating deoxyribose sugars and phosphate groups, providing stability to the molecule.
- Base Pairing: The purine and pyrimidine bases (adenine, thymine, guanine, and cytosine) are located within the interior of the helix. The two strands are held together by hydrogen bonds between complementary bases. Adenine always pairs with thymine through two hydrogen bonds, and guanine always pairs with cytosine through three hydrogen bonds.
- Complementary Base Pair Rule: The base pairing between adenine-thymine and guanine-cytosine is known as the complementary base pair rule. It ensures that the two DNA strands are complementary to each other, meaning that the sequence of bases in one strand determines the sequence in the other.
- Chargaff’s Rules: The base composition of DNA follows Chargaff’s rules. These rules state that the amount of adenine is equal to the amount of thymine (A = T), and the amount of guanine is equal to the amount of cytosine (G = C). Additionally, the sum of purines (A + G) is equal to the sum of pyrimidines (C + T).
- Helix Dimensions: The DNA helix has a diameter of approximately 20 nanometers (20Å). The distance between adjacent nucleotides along the helix axis is approximately 0.34 nanometers (3.4Å). A full turn of the helix spans a length of about 3.4 nanometers (34Å) and consists of approximately 10 base pairs.
- Major and Minor Grooves: The DNA helix exhibits two distinct grooves. The major groove, with a width of approximately 2.2 nanometers (2.2nm), and the minor groove, with a width of approximately 1.2 nanometers (1.2nm). These grooves provide sites for protein binding and interaction with other molecules.
According to Chargaff’s principles (E.E. Chargff, 1950), A = T and G = C; as a corollary, purines (A+G) = ∑ pyrimidines (C+T); similarly, (A+C) = (G+T). It also states that the ratio of (A + T) to (G + C) within a species is constant (range 0.4 to 1.9). DNA has a diameter of 20nm or 20. Along the axis, adjacent bases are separated by 0.34 nm or 3.4. The length of a complete turn of helix is 3.4 nm or 34, which corresponds to 10 base pairs per turn. The DNA helix has a shallow groove known as the minor groove (~1.2nm) and a deep groove known as the major groove (~.2nm).
The features of the Watson-Crick structure of DNA provide the basis for its replication, transcription, and encoding of genetic information. Understanding the unique characteristics of DNA’s double helix is crucial for unraveling its biological functions and processes.
Watson-Crick Model of DNA Summary
The Watson-Crick model of DNA, proposed by James Watson and Francis Crick, provides a comprehensive understanding of the structure and properties of DNA. Here is a summary of the key features of the Watson-Crick model:
- Double Helix: DNA is composed of two helical strands that coil around each other to form a double helix structure resembling a twisted ladder. The two strands are antiparallel, meaning they run in opposite directions.
- 5′ and 3′ Ends: Each DNA strand has a 5′ end and a 3′ end. The 5′ end of one strand is aligned with the 3′ end of the other strand.
- Diameter and Length: The DNA double helix has a uniform diameter of approximately 2 nanometers (nm). Each complete turn of the helix is about 3.6 nm in length, and there are 10.5 base pairs per turn.
- Base Pairing: The nitrogenous bases, Adenine (A), Thymine (T), Guanine (G), and Cytosine (C), are located inside the double helix. Adenine always pairs with Thymine through two hydrogen bonds, while Guanine always pairs with Cytosine through three hydrogen bonds.
- Major and Minor Grooves: The spiral arrangement of the two DNA strands creates distinct grooves. The major groove is larger and the minor groove is smaller. These grooves play a role in DNA-protein interactions and the binding of regulatory proteins.
- Complementary Base Pairs: The base pairing in DNA is highly specific, following the rule of complementary base pairs. A purine base (A or G) always pairs with a pyrimidine base (T or C), maintaining the balance between purines and pyrimidines.
- Hydrogen Bonds: Hydrogen bonds stabilize the base pairing between complementary bases in the double helix. Adenine-thymine pairs form two hydrogen bonds, while guanine-cytosine pairs form three hydrogen bonds.
- Sugar-Phosphate Backbone: The alternating sugar (deoxyribose) and phosphate groups form the backbone of the DNA double helix. The sugar-phosphate backbones are located on the exterior of the helix.
- Chargaff’s Rule: The base composition of DNA follows Chargaff’s rule, which states that the amount of Adenine is equal to Thymine, and the amount of Guanine is equal to Cytosine. This balance between the base pairs is crucial for the stability and proper functioning of DNA.
- Function and Replication: The structure of DNA elucidated by the Watson-Crick model has provided insights into DNA replication, gene expression, and protein synthesis. It serves as a template for DNA replication, transcription, and translation, enabling the transmission and utilization of genetic information in cells.
Overall, the Watson-Crick model of DNA has been instrumental in our understanding of DNA’s structure and function, paving the way for advances in genetics, molecular biology, and various fields of biological research.
Biological Importance of DNA
DNA, or deoxyribonucleic acid, plays a fundamental role in biology and carries significant biological importance. Here are some key aspects of DNA’s biological significance:
- Genetic Information: DNA serves as the repository of genetic information in all living organisms. It contains the instructions that guide the development, growth, and functioning of an organism. The sequence of nucleotides in DNA determines the genetic code, which specifies the synthesis of proteins and governs the traits and characteristics of an organism.
- Inheritance: DNA is responsible for the inheritance of traits from parents to offspring. Through the process of reproduction, DNA is passed from one generation to the next, ensuring the transmission of genetic information. The unique combination of DNA sequences inherited from both parents contributes to the diversity and variability observed within and across species.
- Protein Synthesis: DNA serves as a template for protein synthesis. During the process of transcription, DNA is transcribed into messenger RNA (mRNA), which carries the genetic information to the ribosomes. Subsequently, during translation, the mRNA is used as a template to synthesize proteins, which perform essential functions in cells and organisms.
- Regulation of Gene Expression: DNA plays a crucial role in regulating gene expression, determining when and to what extent genes are turned on or off. Various mechanisms, such as DNA methylation and histone modification, can modify the structure and accessibility of DNA, influencing gene expression patterns. This regulation allows cells to respond to their environment and develop specialized functions.
- Evolution and Species Diversity: DNA underlies the process of evolution and the development of species diversity. Mutations, changes in the DNA sequence, introduce genetic variation, which can be acted upon by natural selection. Over time, these variations can lead to the emergence of new species and the adaptation of organisms to their environments.
- Forensic Identification: DNA analysis has become an invaluable tool in forensic science for identifying individuals. Each person’s DNA is unique, except for identical twins, making it a reliable method for establishing personal identity and determining relationships between individuals.
- Medical Applications: Understanding DNA has led to significant advancements in medicine. DNA sequencing and analysis help diagnose genetic disorders, predict disease susceptibility, and guide personalized treatments. DNA-based technologies, such as recombinant DNA and gene editing techniques like CRISPR-Cas9, hold promise for treating genetic diseases and developing new therapeutic approaches.
- Conservation and Biodiversity: DNA analysis is instrumental in conservation biology and the study of biodiversity. DNA barcoding allows for the identification and classification of species, including those that are endangered or difficult to distinguish based on morphology alone. DNA studies aid in understanding population dynamics, tracking migration patterns, and assessing the genetic health of species and ecosystems.
In summary, DNA’s biological importance lies in its role as the carrier of genetic information, its involvement in protein synthesis and gene regulation, its contribution to evolution and species diversity, its applications in forensics and medicine, and its significance in conservation and biodiversity studies. Understanding DNA has revolutionized our understanding of life and has far-reaching implications in various fields of science and technology.
How DNA strands are connected to each other?
The two strands of DNA are connected to each other through hydrogen bonds and a backbone of sugar-phosphate molecules.
The hydrogen bonds form between the nitrogenous bases of the two DNA strands. Adenine (A) always pairs with Thymine (T) through two hydrogen bonds, and Guanine (G) always pairs with Cytosine (C) through three hydrogen bonds. These specific base pairings ensure the complementary nature of the DNA strands.
The sugar-phosphate backbone runs along the outside of the double helix. It is composed of alternating deoxyribose sugar molecules and phosphate groups. The sugar molecules are linked together by phosphodiester bonds, which connect the 3′ carbon of one sugar to the 5′ carbon of the next sugar in the chain. This linkage creates a repeating sugar-phosphate backbone that provides stability and structure to the DNA molecule.
The combination of hydrogen bonds between the nitrogenous bases and the sugar-phosphate backbone holding the strands together forms the double helix structure of DNA. This connection allows for the precise pairing of the complementary bases on each strand and maintains the integrity of the genetic information encoded in the DNA molecule.
Chargaff’s rule
Chargaff’s rule, proposed by Erwin Chargaff, states that in a double-stranded DNA molecule, the amount of adenine (A) is equal to the amount of thymine (T), and the amount of guanine (G) is equal to the amount of cytosine (C). This rule can be summarized as A = T and G = C.
Chargaff’s rule is based on the observation that the composition of DNA varies between different species. By analyzing the base composition of DNA from different organisms, Chargaff discovered that the percentages of A and T are always approximately equal, as are the percentages of G and C. This finding was a crucial clue in understanding the structure and function of DNA.
The complementary base pairing in DNA, where A pairs with T and G pairs with C, is consistent with Chargaff’s rule. The pairing of these bases through hydrogen bonds ensures that the amount of purines (A and G) is always equal to the amount of pyrimidines (T and C), thus maintaining the structural integrity and stability of the DNA molecule.
Chargaff’s rule played a significant role in the discovery of the double helical structure of DNA by Watson and Crick. It provided valuable insight into the fundamental principles of DNA composition and helped establish the foundation for our understanding of the genetic code and DNA replication.
What are the key features of the Watson and Crick DNA model?
The key features of the Watson and Crick DNA model are as follows:
- Double Helix Structure: The model proposed by Watson and Crick describes DNA as a double-stranded helix. The two strands are coiled around each other in a twisted ladder-like structure.
- Antiparallel Strands: The two strands in the DNA molecule run in opposite directions. One strand has a 5′ end (with a phosphate group) and a 3′ end (with a hydroxyl group), while the other strand has a 3′ end and a 5′ end. This arrangement is known as antiparallel orientation.
- Complementary Base Pairing: The DNA strands are held together by complementary base pairing. Adenine (A) always pairs with thymine (T) through two hydrogen bonds, and guanine (G) always pairs with cytosine (C) through three hydrogen bonds. This pairing ensures the stability and integrity of the DNA molecule.
- Base Pairing Rule: The base pairing rule in the Watson and Crick model states that the number of adenine (A) bases is equal to the number of thymine (T) bases, and the number of guanine (G) bases is equal to the number of cytosine (C) bases. This rule is also known as Chargaff’s rule.
- Sugar-Phosphate Backbone: The outside of the DNA double helix is formed by alternating sugar (deoxyribose) and phosphate groups. These sugar-phosphate backbones provide structural support and stability to the DNA molecule.
- Major and Minor Grooves: The twisting of the DNA double helix creates two grooves along its length. The major groove is wide and the minor groove is narrow. These grooves provide binding sites for proteins and other molecules involved in various cellular processes.
- Replication and Genetic Information: The Watson and Crick model of DNA provided insights into how DNA replicates and carries genetic information. The complementary base pairing allows for accurate replication, with each strand serving as a template for the synthesis of a new complementary strand.
- Molecular Basis of Inheritance: The discovery of the double helix structure of DNA by Watson and Crick laid the foundation for understanding the molecular basis of inheritance. It revealed how genetic information is encoded and transmitted in the form of DNA sequences.
These key features of the Watson and Crick DNA model have had a profound impact on our understanding of genetics, molecular biology, and the central role of DNA in life processes.
How did Chargaff’s rules contribute to the development of the Watson and Crick model?
Chargaff’s rules played a crucial role in the development of the Watson and Crick model of DNA. Erwin Chargaff, a biochemist, discovered certain regularities in the composition of DNA bases, which provided important insights for understanding the structure of DNA. The contributions of Chargaff’s rules to the Watson and Crick model are as follows:
- Base Pairing: Chargaff’s rules stated that in DNA, the amount of adenine (A) is equal to the amount of thymine (T), and the amount of guanine (G) is equal to the amount of cytosine (C). This observation indicated a specific pairing between the bases, suggesting that A always pairs with T, and G always pairs with C.
- Complementary Base Pairing: Chargaff’s rules provided evidence for the concept of complementary base pairing in DNA. The equal ratios of A to T and G to C indicated that the bases are paired in a specific manner. This information was crucial for Watson and Crick in proposing their double helix model, where the complementary pairing of bases stabilizes the structure.
- Structural Constraints: Chargaff’s rules also implied that the total purine (A + G) content is equal to the total pyrimidine (T + C) content in DNA. This suggested that the DNA molecule had a consistent and uniform structure. Watson and Crick used this information to develop their model, ensuring that the double helix structure maintained equal spacing between the bases.
By following Chargaff’s rules, Watson and Crick were able to propose a model that incorporated the principles of complementary base pairing and the equal ratios of base pairs. Chargaff’s rules served as a guiding principle for understanding the structure and properties of DNA, and their contribution was instrumental in the development of the Watson and Crick model.
What was the significance of Watson and Crick’s discovery of the double helix structure?
The discovery of the double helix structure by James Watson and Francis Crick in 1953 was a groundbreaking milestone in the field of molecular biology with immense significance. The key significance of their discovery can be summarized as follows:
- Understanding DNA Structure: Watson and Crick’s discovery provided the first accurate and comprehensive understanding of the three-dimensional structure of DNA. They proposed that DNA consists of two complementary strands that are twisted around each other in a double helix configuration. This knowledge laid the foundation for further research and exploration of the genetic material.
- Genetic Information Storage: The double helix structure revealed how genetic information is stored and transmitted in living organisms. The structure showed that the sequence of nucleotide bases along the DNA strands carries the genetic code, which determines the synthesis of proteins and other cellular functions. It established the connection between DNA structure and the hereditary characteristics of organisms.
- Base Pairing and Replication: Watson and Crick’s model demonstrated the specific base pairing of adenine (A) with thymine (T) and guanine (G) with cytosine (C). This base pairing provided insights into DNA replication, as it explained how each strand could serve as a template for the synthesis of a new complementary strand during cell division.
- Molecular Interactions: The discovery of the double helix structure highlighted the molecular interactions that stabilize the DNA molecule. It revealed that hydrogen bonds form between the base pairs, holding the two strands together. This understanding of molecular interactions in DNA paved the way for studying other biological macromolecules and their functions.
- Advances in Genetics and Biotechnology: The knowledge gained from the double helix structure has revolutionized the fields of genetics and biotechnology. It has enabled scientists to unravel the mechanisms of inheritance, gene expression, and genetic engineering. It has also facilitated advancements in DNA sequencing, genetic diagnostics, and the development of novel therapies and treatments.
- Impact on Scientific Community: Watson and Crick’s discovery sparked a scientific revolution and transformed our understanding of life at the molecular level. It inspired generations of scientists and paved the way for further discoveries in molecular biology, genetics, and related disciplines.
Overall, the significance of Watson and Crick’s discovery of the double helix structure lies in its profound impact on our understanding of DNA, genetics, and the fundamental mechanisms of life. It laid the groundwork for countless scientific breakthroughs and continues to shape research in biological sciences to this day.
How does the Watson and Crick model explain the process of DNA replication?
The Watson and Crick model of DNA, also known as the double helix structure, provides a key explanation for the process of DNA replication. According to this model, DNA replication occurs through a semi-conservative mechanism, where each original DNA strand serves as a template for the synthesis of a new complementary strand. The process can be summarized as follows:
- Unwinding: The double-stranded DNA molecule unwinds and separates into two individual strands. This unwinding is facilitated by enzymes called DNA helicases, which break the hydrogen bonds between the base pairs and create a replication fork.
- Complementary Base Pairing: Each separated DNA strand serves as a template for the synthesis of a new complementary strand. The existing bases on the template strand determine the sequence of the new strand. Adenine (A) on the template strand pairs with thymine (T) and guanine (G) pairs with cytosine (C) through specific hydrogen bonding.
- DNA Polymerization: Enzymes known as DNA polymerases catalyze the addition of nucleotides to the growing DNA strand. DNA polymerase recognizes the exposed bases on the template strand and adds complementary nucleotides one by one, following the base-pairing rules. The nucleotides are joined together by phosphodiester bonds, forming the sugar-phosphate backbone of the new strand.
- Leading and Lagging Strands: DNA replication occurs in a continuous manner on one of the template strands, known as the leading strand. The DNA polymerase synthesizes the new complementary strand continuously in the 5′ to 3′ direction, following the replication fork. On the other template strand, called the lagging strand, replication occurs discontinuously in small fragments called Okazaki fragments. The fragments are later joined together by DNA ligase.
- Replication Fork Movement: As DNA replication progresses, the replication fork moves along the DNA molecule. The unwinding and synthesis of new strands occur simultaneously at the replication fork. Multiple replication forks can be present on a DNA molecule to expedite the replication process.
- Proofreading and Repair: DNA polymerases have proofreading capabilities to ensure the accuracy of replication. They can detect and correct errors in base pairing. Additionally, various repair mechanisms exist to fix any mistakes or damaged regions in the replicated DNA.
By following the Watson and Crick model, DNA replication ensures the faithful transmission of genetic information from one generation to the next. The model explains how each strand of the original DNA molecule serves as a template for the synthesis of new strands, resulting in two identical copies of the DNA molecule.
What is the role of complementary base pairing in the Watson and Crick DNA model?
In the Watson and Crick DNA model, complementary base pairing plays a crucial role in the structure and replication of DNA. The model describes the double helix structure of DNA, where two strands of nucleotides are twisted around each other in a ladder-like arrangement.
Complementary base pairing refers to the specific pairing of nucleotide bases between the two DNA strands. In DNA, there are four nucleotide bases: adenine (A), thymine (T), cytosine (C), and guanine (G). The complementary base pairs are A with T and C with G. This means that adenine always pairs with thymine, and cytosine always pairs with guanine.
The base pairing is based on hydrogen bonding between the bases. Adenine forms two hydrogen bonds with thymine, while cytosine forms three hydrogen bonds with guanine. These hydrogen bonds provide stability to the DNA structure and hold the two strands together.
The complementary base pairing is significant because it allows for accurate DNA replication and the transmission of genetic information. During DNA replication, the two strands of the double helix separate, and each strand serves as a template for the synthesis of a new complementary strand. The base pairing rules ensure that the new strands are complementary to the original strands, resulting in two identical copies of the DNA molecule.
Additionally, the complementary base pairing plays a crucial role in gene expression and protein synthesis. The sequence of bases in DNA forms the genetic code, where each three-base sequence, called a codon, codes for a specific amino acid. The accurate base pairing ensures that the correct sequence of amino acids is encoded in the DNA and subsequently translated into proteins.
In summary, complementary base pairing in the Watson and Crick DNA model ensures the stability of the double helix structure, enables accurate DNA replication, and allows for the faithful transmission of genetic information.
What evidence or data did Watson and Crick use to support their model of DNA?
Watson and Crick’s model of DNA was primarily based on a combination of existing experimental data and theoretical insights from other scientists. Although they did not conduct new experiments themselves, they incorporated crucial information from various sources to formulate their model. Some of the key evidence and data that contributed to their understanding of DNA are:
- X-ray crystallography by Rosalind Franklin and Maurice Wilkins: Franklin and Wilkins used X-ray crystallography techniques to study the structure of DNA fibers. Franklin’s X-ray diffraction images provided critical information about the helical nature of DNA and the spacing of its repeating units. Watson and Crick had access to Franklin’s unpublished data and utilized it to refine their model.
- Chargaff’s rules: Erwin Chargaff, a biochemist, discovered that in DNA, the amount of adenine (A) is equal to the amount of thymine (T), and the amount of cytosine (C) is equal to the amount of guanine (G). This observation, known as Chargaff’s rules, indicated a consistent pairing between the bases and provided hints about the structure of DNA.
- Linus Pauling’s alpha-helix structure: Linus Pauling, a prominent chemist, had proposed a helical structure for proteins based on their repeating patterns. Although Pauling’s alpha-helix structure was ultimately not applicable to DNA, it influenced Watson and Crick to consider a helical structure for DNA.
- Knowledge of chemical bonding: Watson and Crick had a strong background in chemistry, which helped them understand the principles of chemical bonding. They knew that hydrogen bonds were relatively weak but crucial for maintaining the stability of biological molecules. Applying this knowledge, they hypothesized that hydrogen bonding between specific base pairs could explain the complementarity of DNA strands.
By combining these pieces of information, Watson and Crick deduced the correct structure of DNA—a double helix with antiparallel strands held together by complementary base pairing. Their model elegantly explained how DNA could store and transmit genetic information, and it provided a solid foundation for understanding DNA replication and the central dogma of molecular biology.
How did the Watson and Crick model of DNA revolutionize our understanding of genetics and molecular biology?
The Watson and Crick model of DNA, proposed in 1953, revolutionized our understanding of genetics and molecular biology in several significant ways. Here are some key impacts of their model:
- Structure of DNA: Watson and Crick’s model revealed the double helix structure of DNA, which provided a physical framework for understanding how genetic information is stored and transmitted. It explained how the complementary base pairing allows for the accurate replication of DNA during cell division.
- DNA as the Genetic Material: The discovery of the DNA structure solidified the understanding that DNA is the molecule responsible for carrying genetic information. Prior to this, there were debates about whether proteins or nucleic acids were the carriers of genetic traits. The Watson and Crick model supported the notion that DNA, with its specific base sequences, encodes the instructions for protein synthesis and hereditary traits.
- Genetic Code and Protein Synthesis: The model’s base pairing rules and the understanding of the genetic code paved the way for deciphering how DNA sequences are translated into proteins. The codon-anticodon correspondence explained how specific combinations of three DNA bases (codons) specify the synthesis of particular amino acids during protein synthesis. This formed the basis for our understanding of gene expression and the central dogma of molecular biology.
- Heredity and Evolution: The Watson and Crick model provided insights into the mechanisms of heredity and evolution. It explained how DNA replication ensures the faithful transmission of genetic information from one generation to the next. It also shed light on the potential for genetic variations through mutations and recombination, which are the basis for genetic diversity and the driving forces behind evolution.
- Advances in Molecular Biology: The discovery of the DNA structure stimulated significant advancements in molecular biology. It opened doors for further research into DNA replication, repair, and recombination, as well as the regulation of gene expression. It also facilitated the development of techniques such as polymerase chain reaction (PCR), DNA sequencing, and genetic engineering, which have revolutionized various fields including medicine, agriculture, and biotechnology.
Overall, the Watson and Crick model of DNA provided a fundamental understanding of the molecular basis of genetics. It laid the groundwork for numerous breakthroughs in molecular biology, enabling us to unravel the mechanisms underlying inheritance, gene expression, and the intricate workings of living organisms.
Here are 10 recent research findings on DNA models
- DNA can be used to create new materials. Scientists have been able to use DNA to create new materials with properties that are not found in naturally occurring materials. For example, DNA can be used to create materials that are transparent, conductive, and even self-healing.
- DNA can be used to create new drugs. Scientists are developing new drugs that use DNA as a delivery mechanism. These drugs can target specific cells or tissues, which could make them more effective and less harmful than traditional drugs.
- DNA can be used to create new sensors. Scientists are developing new sensors that use DNA to detect specific molecules. These sensors could be used to diagnose diseases, monitor environmental pollutants, and even track the movement of cells in the body.
- DNA can be used to create new biofuels. Scientists are developing new biofuels that are made from DNA. These biofuels could be used to replace fossil fuels, which would help to reduce greenhouse gas emissions.
- DNA can be used to create new batteries. Scientists are developing new batteries that are made from DNA. These batteries could be used to power devices that are too small for traditional batteries, such as implantable medical devices and wearable electronics.
- DNA can be used to create new electronics. Scientists are developing new electronics that are made from DNA. These electronics could be used to create new types of computers, sensors, and other devices.
- DNA can be used to create new DNA editing tools. Scientists are developing new DNA editing tools that use CRISPR-Cas9 technology. These tools could be used to correct genetic defects, create new crops and livestock, and even develop new forms of gene therapy.
- DNA can be used to create new vaccines. Scientists are developing new vaccines that use DNA as a delivery mechanism. These vaccines could be more effective and less expensive than traditional vaccines.
- DNA can be used to create new cancer treatments. Scientists are developing new cancer treatments that use DNA as a delivery mechanism. These treatments could target cancer cells more effectively and with fewer side effects than traditional treatments.
- DNA can be used to create new personalized medicine. Scientists are developing new personalized medicine approaches that use DNA to tailor treatments to individual patients. This could lead to more effective and less expensive treatments for a variety of diseases.
FAQ
What model of DNA was discovered by Watson and Crick?
Watson and Crick discovered the double helix model of DNA.
What did Watson and Crick’s model of DNA show?
Watson and Crick’s model of DNA showed that DNA is a double-stranded molecule made up of two helical chains spirally coiled around a common axis.
Which description best explains Watson and Crick’s DNA model?
Watson and Crick’s DNA model can be described as a right-handed double helix structure with two antiparallel strands.
How to draw Watson and Crick’s model of DNA?
Drawing Watson and Crick’s model of DNA involves representing two strands coiled in a helical shape with complementary base pairing (adenine with thymine, and guanine with cytosine) and showing the sugar-phosphate backbones on the outside.
How did Chargaff’s rules help Watson and Crick’s model of DNA?
Chargaff’s rules helped Watson and Crick’s model of DNA by providing important insights into the base composition. Chargaff’s rule states that the amount of adenine is equal to thymine, and the amount of guanine is equal to cytosine, which guided Watson and Crick in understanding the complementary base pairing.
How did Watson and Crick develop the model of DNA?
Watson and Crick developed their model of DNA through a combination of scientific data, analysis of X-ray crystallography images (such as those taken by Rosalind Franklin), and model building based on chemical principles.
What did Watson and Crick’s model of DNA look like?
Watson and Crick’s model of DNA featured a double helix structure with two strands, a uniform diameter of approximately 2 nanometers, and a repeating pattern of base pairs along the length.
What does Watson and Crick’s model of DNA demonstrate?
Watson and Crick’s model of DNA demonstrated the complementary base pairing between adenine and thymine, and between guanine and cytosine, forming the steps of the DNA ladder.
When did Watson and Crick present their DNA model?
Watson and Crick presented their DNA model on April 25, 1953, in a paper published in the scientific journal Nature titled “Molecular Structure of Nucleic Acids: A Structure for Deoxyribose Nucleic Acid.”
Why was Watson and Crick’s first DNA model incorrect?
The first DNA model proposed by Watson and Crick had some inaccuracies and did not fully represent the precise structure of DNA. However, through continued research and collaboration, they refined their model to accurately depict the double helix structure.
References
- https://embryo.asu.edu/pages/genetical-implications-structure-deoxyribonucleic-acid-1953-james-watson-and-francis-crick
- https://www.nature.com/scitable/topicpage/discovery-of-dna-structure-and-function-watson-397/
- https://www.khanacademy.org/humanities/big-history-project/life/knowing-life/a/crick-watson-and-franklin2
- https://profiles.nlm.nih.gov/spotlight/sc/feature/doublehelix
- https://www.khanacademy.org/science/biology/dna-as-the-genetic-material/dna-discovery-and-structure/a/discovery-of-the-structure-of-dna
- https://thesciencenotes.com/watson-and-crick-double-helix-model-of-dna/
- https://unacademy.com/content/nta-ugc/study-material/pharmaceutical-analysis/the-watson-crick-structure-of-dna/
- https://ib.bioninja.com.au/standard-level/topic-2-molecular-biology/26-structure-of-dna-and-rna/watson–crick.html
- http://www.pbs.org/wgbh/aso/databank/entries/do53dn.html
- https://geneticeducation.co.in/dna-deoxyribonucleic-acid-definition-structure-function-evidence-and-types/
- https://www.mun.ca/biology/scarr/Watson-Crick_Model.html
- https://www.toppr.com/ask/question/explain-watson-and-cricks-model-of-dna/
- https://microbenotes.com/watson-and-crick-dna-model/
- https://unacademy.com/content/nta-ugc/study-material/pharmaceutical-analysis/the-watson-crick-structure-of-dna/
- https://byjus.com/neet/double-helix-structure-of-dna/
- https://www.merriam-webster.com/dictionary/Watson-Crick%20model
- https://www.vedantu.com/question-answer/explain-watson-and-crick-model-of-dna-class-12-biology-cbse-5f74594c0fd0025e7767ce48
- https://www.sciencehistory.org/historical-profile/james-watson-francis-crick-maurice-wilkins-and-rosalind-franklin
- https://study.com/academy/lesson/watson-crick-model-of-dna.html
- https://biologyreader.com/watson-and-crick-model-of-dna.html
- https://www.history.com/this-day-in-history/watson-and-crick-discover-chemical-structure-of-dna
- https://www.toppr.com/ask/en-us/question/explain-watson-and-cricks-model-of-dna/
- https://royalsociety.org/blog/2018/04/history-of-the-double-helix/
- https://easyhsc.com.au/home-easyhsc/easybio/heredity/cell-replication/watson-crick-dna-model/
- https://www.yourgenome.org/stories/unravelling-the-double-helix/
- http://www.lscollege.ac.in/sites/default/files/e-content/Structure%20of%20DNA.pdf
- https://jackwestin.com/resources/mcat-content/nucleic-acid-structure-and-function/deoxyribonucleic-acid-dna-double-helix-watson-crick-model-of-dna-structure
- https://www.javatpoint.com/dna-structure
- https://annex.exploratorium.edu/origins/coldspring/printit.html
- https://www.nobelprize.org/prizes/medicine/1962/speedread/
- https://dosequis.colorado.edu/Courses/MethodsLogic/papers/WatsonCrick1953.pdf
- https://biologyreader.com/watson-and-crick-model-of-dna.html
- https://www.vedantu.com/question-answer/explain-watson-and-crick-model-of-dna-class-12-biology-cbse-5f74594c0fd0025e7767ce48
- https://undsci.berkeley.edu/the-structure-of-dna-cooperation-and-competition/a-false-start/
- https://www.zigya.com/study/book?class=11&board=cbse&subject=Biology&book=Biology&chapter=Biomolecules&q_type=&q_topic=Nature%20Of%20Bond%20Linking%20Monomers%20In%20A%20Polymer&question_id=BIEN11007981
- https://www.pbs.org/wgbh/evolution/library/06/3/l_063_01.html
- http://www.pbs.org/wgbh/aso/databank/entries/do53dn.html
- https://www.theguardian.com/science/2015/jun/23/sexism-in-science-did-watson-and-crick-really-steal-rosalind-franklins-data
- https://ib.bioninja.com.au/standard-level/topic-2-molecular-biology/26-structure-of-dna-and-rna/watson–crick.html
- https://www.biologydiscussion.com/dna/watson-and-cricks-model-of-double-helix-of-dna-biochemistry/65076
- https://aklectures.com/lecture/structure-and-function/watson-crick-model-of-dna
- https://collection.sciencemuseumgroup.org.uk/objects/co146411/crick-and-watsons-dna-molecular-model-molecular-model
- https://www.khanacademy.org/science/biology/dna-as-the-genetic-material/dna-discovery-and-structure/a/discovery-of-the-structure-of-dna
- https://www.khanacademy.org/humanities/big-history-project/life/knowing-life/a/crick-watson-and-franklin2
