Table of Contents
- DNA replication is a prerequisite for cell division in both prokaryotic and eukaryotic organisms.
- DNA replication is the biological process whereby two identical copies of DNA are synthesised from a single DNA molecule.
- DNA replication guarantees that each daughter cell inherits an identical set of genetic information from its parent cells. DNA polymerases are enzymes responsible for replicating genetic material.
- In both prokaryotic and eukaryotic DNA replications, one old and one new strand are present in the daughter cell, making them semi-conservative DNA replications.
- DNA replication is very similar in prokaryotes and eukaryotes; however, there may be subtle differences due to the size and complexity of the genetic material.
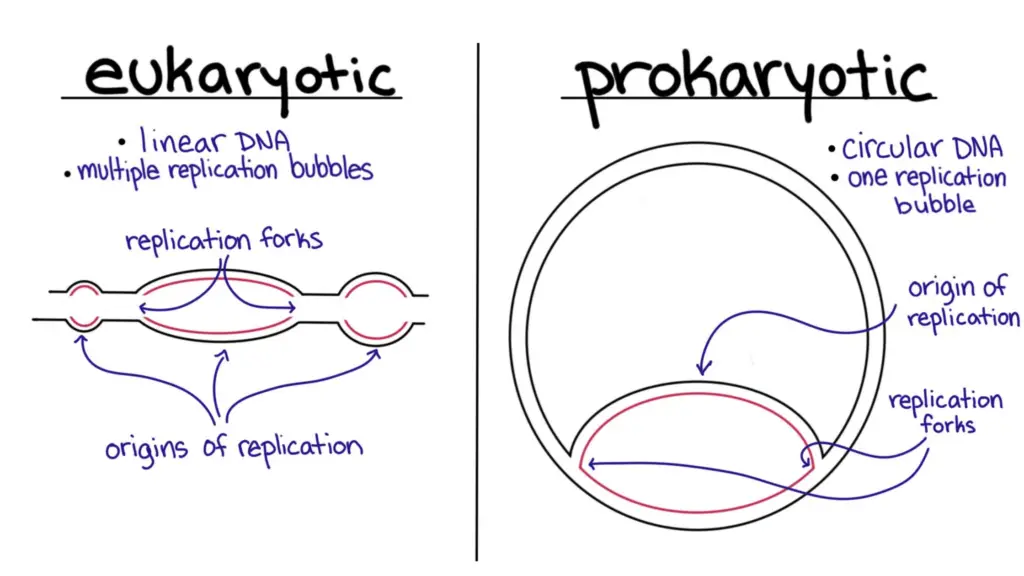
- DNA replication in prokaryotes takes place at a single origin of replication, whereas in eukaryotes it takes place at multiple origins of replication.
Prokaryotic DNA Replication
- DNA replication in prokaryotes, such as bacteria and archaea, is the process by which the genome is copied so that a daughter cell can be created.
- The cytoplasm of prokaryotes contains a circular molecule of DNA with two strands of DNA. It is important to note that the DNA of prokaryotes only has a single replication origin.
- DNA helicase, by severing hydrogen bonds between the nucleic acid’s nitrogenous bases, unwinds the DNA at the replication origin.
- The Y-shaped structure so formed is known as a replication fork.
- Due to the presence of just one replication origin in bacterial DNA, only two replication forks are generated during replication.
- Both of these replication forks can move in either direction. The two unwound strands that will be used as replication templates are stabilised by single-strand DNA-binding (SSB) proteins.
- Using the template strand as a template, the enzyme RNA primase creates a five to ten base pair long RNA primer.
- DNA polymerases I, II, and III are all essential for DNA replication in prokaryotes.
- DNA polymerase III is responsible for both the initiation and the elongation phases of DNA replication in prokaryotes.
- Nucleotides are incorporated by DNA polymerase III from the 5′ end to the 3′ end. The DNA double helix is antiparallel, thus one strand travels in a 5′ to 3′ orientation (leading strand).
- The 3–5 direction is the direction of the opposite strand (lagging strand). Okazaki fragments are constantly being generated because the lagging strand requires RNA primers in order to synthesis DNA in the 5′ to 3′ orientation.
- DNA polymerases I and II are responsible for repairing DNA and filling in gaps.
- DNA polymerase I is responsible for eliminating the RNA primer.
Eukaryotic DNA Replication
- Before a cell divides, the eukaryotic genome undergoes a process called DNA replication.
- Although eukaryotic and prokaryotic DNA replication share a similar underlying mechanism, there are notable distinctions due to the larger size and more complex structure of eukaryotic DNA.
- All eukaryotic DNA molecules are double-stranded and linear.
- DNA in eukaryotes is roughly 50 times as abundant as DNA in bacteria.
- Furthermore, histones bundle eukaryotic DNA firmly into the nucleus of the cell.
- There are three stages of DNA replication: initiation, elongation, and termination.
Initiation
- Multiple replication sources are used in the replication of eukaryotic DNA.
- In a chromosome with numerous replication sources, various bubbles of replication will emerge.
- DNA helicase and SSBs work together at both replication origins to unwind and stabilise the two templates.
Elongation
- DNA polymerases add additional nucleotides to the 3′ ends of preexisting strands during the elongation process.
- Eukaryotic DNA replication requires the actions of three distinct DNA polymerases: DNA polymerase,, and.
- DNA replication is started by DNA polymerase, whereas DNA polymerases and participate in replication elongation.
- DNA polymerase, like DNA polymerase, needs an RNA primer to synthesise the new DNA strand and then removes the primer after synthesis is complete.
- Similar to prokaryotic DNA replication, both the leading and lagging strands are generated.
Termination
- Replication stops when the leading strand of one replication bubble collides with the trailing strand of another replication bubble.
- After that, the RNA primer is taken out of the way, and the free-floating DNA polymerases fill in the resulting void.
- DNA ligase joins the cuts in the DNA.
Similarities Between Prokaryotic and Eukaryotic DNA Replication
- Before the nuclear division in both prokaryotes and eukaryotes, DNA replication takes place.
- DNA replication occurs on double-stranded DNA in both prokaryotic and eukaryotic cells.
- DNA helicase is responsible for unwinding the DNA of both prokaryotic and eukaryotic cells.
- Proteins that bind just to single strands of DNA help keep the DNA in its unravelled state (SSB).
- DNA replication is a multistep process that occurs in both prokaryotic and eukaryotic cells and is carried out by an enzyme complex called DNA polymerases.
- DNA polymerases of all types copy DNA from 5′ to 3′.
- Primers made of RNA are essential for starting both strands of DNA replication from scratch.
- The enzyme known as primase is responsible for synthesising the RNA primer.
- Both bacterial and eukaryotic DNA replications are only half-conservative, resulting in the presence of one parent strand and one daughter strand in the progeny.
- Since DNA replication can proceed in either direction, it is safe to say that both prokaryotic and eukaryotic cells replicate their DNA in two directions.
- Both methods of DNA replication result in the formation of leading and lagging strands.
- The lagging strand generates Okazaki fragments, which are short pieces of DNA that are recombined.
- Both methods of replication take around an hour to complete.
difference between prokaryotic and eukaryotic dna replication
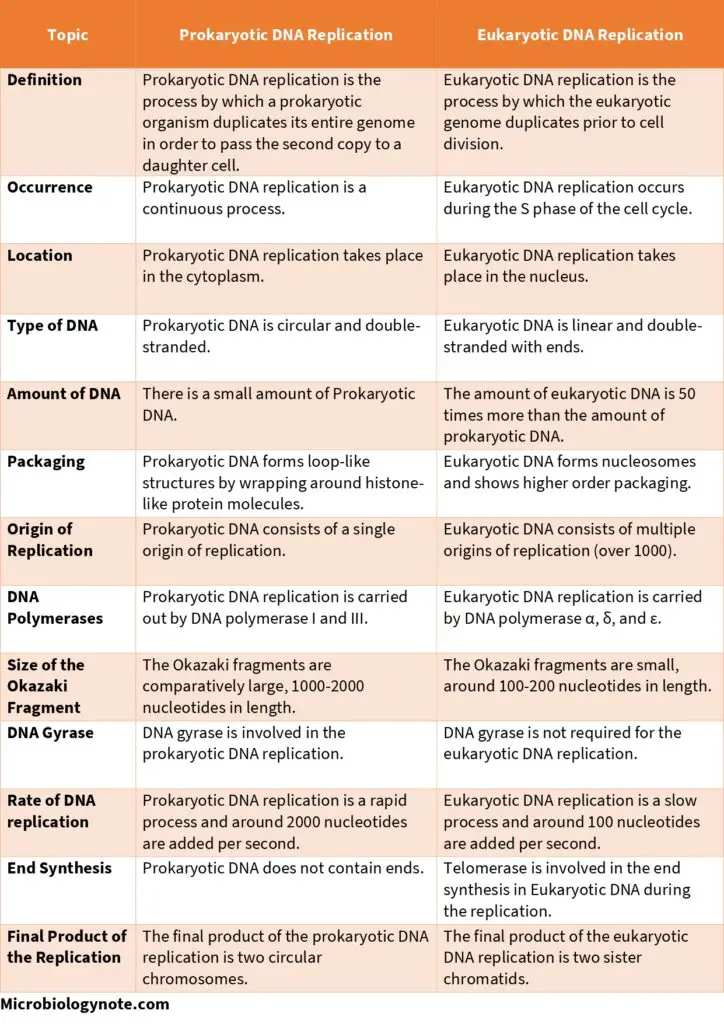
1. Location
- The nucleus and other organelles that are surrounded by membranes, including as the mitochondria, endoplasmic reticulum, and golgi bodies, are absent in prokaryotes. In prokaryotes, DNA is found in a DNA-protein complex known as nucleoid. The cellular cytoplasm is the site of the replication.
- eukaryotes, which have a nucleus surrounded by a membrane, keep their DNA safe there. Because of this, DNA replication in eukaryotes takes place in the nucleus.
2. Stage of Cell Division
- Binary fission or budding, the two most common methods of cell division in prokaryotes, occurs immediately after DNA replication.
- DNA replication occurs during the synthesis (S) phase of the cell cycle, which is a somewhat complex process in eukaryotes.
3. Initiation
- Replication of DNA begins at what is called an origin of replication and concludes at what are also called termination sites. A replication unit, or replicon, is the segment of DNA that contains these two locations.
- The DNA of prokaryotes is packaged into chromosomes, which are in turn packaged into smaller circular DNA molecules called plasmids. There is only one replication origin and one replicon per DNA molecule in prokaryotes. In addition, these genesis sites tend to be longer than their eukaryotic counterparts.
- The eukaryotic genome is relatively big and is structured into linear chromosomes. Each chromosome has several origins of replication due to the large amount of data that must be replicated. It is possible for DNA replication to begin at many origins and stop at their respective termination sites. That’s why each chromosome has several replicons, which allows DNA replication to proceed at a much quicker rate. The entire human genome, which consists of around 3.2 billion base pairs, can be copied in less than an hour. It would take a month to replicate a single chromosome if DNA replication required a single replicon.
4. Direction of Replication
- DNA replication assembly progresses along the DNA molecule once it has been commenced; the replication fork indicates the precise location of replication. Both prokaryotes and eukaryotes replicate their DNA in two different orientations, starting at the replication origin.
- It has been revealed, however, that DNA replication can occur in only one way in some plasmids found in bacterial cells. With this type of plasmid replication, numerous linear copies of the circular DNA are generated and subsequently circularised.
5. Enzymes
DNA replication in prokaryotes and eukaryotes utilises comparable enzymes, although DNA replication in eukaryotes is more intricate and diverse. DNA replication initiator proteins such as single-stranded DNA-binding protein (SSB), primase, DNA helicase, and DNA ligase are shared between prokaryotes and eukaryotes.
DNA polymerase, which plays a role in the replication of mitochondrial DNA, is also present in eukaryotic organisms. There is also variation in the activity of topoisomerases, enzymes that control the winding and unwinding of DNA as the replication fork moves. DNA gyrase, a type II topoisomerase, is commonly used by prokaryotes to create a nick in both DNA strands. As the replication fork advances, most eukaryotes use single-strand-cutting type I topoisomerases.
6. Okazaki fragments
- DNA replication involves the continuous synthesis of one strand and the discontinuous, fragmentary synthesis of the complementary strand. For convenience, we will refer to the former strand as the leading strand, the later as the lagging strand, and the intermediate pieces as the Okazaki fragments. The discrepancy arises because DNA strands are antiparallel, but DNA polymerase acts in only one direction.
- The usual size of an Okazaki fragment in the prokaryotic organism Escherichia coli (E. coli) is between 1000 and 2000 nucleotides.
- Okazaki fragments in eukaryotes are typically between 100 and 200 nucleotides in length. They are formed at a slower rate than that seen in prokaryotes, although being shorter in length.
7. Termination
- DNA replication in both prokaryotes and eukaryotes is halted at predetermined places called termination points.
- Midway around the circular chromosome in prokaryotes is a single termination site. As the two branches of replication converge here, the process is halted.
- Different origins of replication are located at different points along the chromosome in eukaryotes, which results in linear DNA molecules with multiple termination sites. An unusual end-replication problem, in which some DNA at the chromosomal ends does not get copied, is characteristic to eukaryotic DNA replication. As a result, the trailing strand is noticeably shorter than the leading one. In eukaryotes, telomeres, which are made up of non-coding, repetitive DNA sequence, are present at the ends of chromosomes to combat this issue.
8. Rate of DNA replication
- In a prokaryote, around 2000 nucleotides are added to the DNA strand every second, making DNA replication a very fast process.
- The addition of just about 100 nucleotides to the DNA of a eukaryotic cell every second makes DNA replication a very sluggish process.
9. Packaging
- Prokaryotic DNA Replication: DNA wraps around histone-like protein molecules to form loop-like structures.
- Higher order packaging and the formation of nucleosomes characterise the replication of eukaryotic DNA.
10. Amount of DNA
- It is possible to replicate a little quantity of prokaryotic DNA.
- Replication of eukaryotic DNA: eukaryotic genomes contain 50 times as much DNA as prokaryotic genomes.
DNA replication protein list
| Function | Bacteria | Eukaryotes |
| Initiator; binds origin | Dna A | ORC |
| Helicase loader | DnaC | Cdc6 and Cdt1 |
| Helicase | DnaB | MCM complex |
| Single-strand binding | SSB proteins | RPA |
| Primer synthesis | DnaG primase | Pol α-primase1 |
| Replicative DNA polymerase | DNA polymerase III (C-family polymerase) | DNA polymerase δ and DNA polymerase ε (B-family polymerases) |
| Clamp loader | ϒ complex | RF-C |
| Clamp | β clamp | PCNA |
| Ligase | DNA ligase2 | DNA ligase 12 |
| Primer removal | Ribonuclease H; DNA polymerase I | RNaseH/Fen1 |
References
- https://biologywise.com/prokaryotic-vs-eukaryotic-dna-replication
- https://www.vedantu.com/biology/difference-between-prokaryotic-and-eukaryotic-dna-replication
- https://pediaa.com/difference-between-prokaryotic-and-eukaryotic-dna-replication/
- https://thebiotechnotes.com/2019/10/01/difference-between-prokaryotic-and-eukaryotic-replication/
- https://www.researchgate.net/profile/Lakna-Panawala/publication/316617219_Home_Science_Biology_Molecular_Biology_Difference_Between_Prokaryotic_and_Eukaryotic_DNA_Difference_Between_Prokaryotic_and_Eukaryotic_DNA/links/5907eb77a6fdccd580dd06b0/Home-Science-Biology-Molecular-Biology-Difference-Between-Prokaryotic-and-Eukaryotic-DNA-Difference-Between-Prokaryotic-and-Eukaryotic-DNA.pdf
- https://www.majordifferences.com/2013/03/difference-between-prokaryotic-and.html
- https://www.differencebetween.com/difference-between-prokaryotic-and-vs-eukaryotic-dna/
- https://en.wikipedia.org/wiki/Eukaryotic_DNA_replication#Comparisons_between_prokaryotic_and_eukaryotic_DNA_replication
