Table of Contents
What is a Genetic Code?
The genetic code is a set of rules that living cells use to decipher the information encoded in genetic material (DNA or mRNA sequences). The ribosomes are responsible for carrying out the translation process. Using tRNA (transfer RNA) molecules to carry amino acids and to read the mRNA three nucleotides at a time, they link the amino acids in an mRNA-specified (messenger RNA) order.
- As DNA is a genetic substance, it transmits genetic information from one cell to the next and from one generation to the next.
- At this point, it will be attempted to determine how genetic information is stored within the DNA molecule. On the DNA molecule, are they written in an articulated or encoded language? In the language of codes, what is the genetic code’s nature?
- A DNA molecule contains three types of moieties: phosphoric acid, deoxyribose sugar, and nitrogen bases.
- The genetic information may be encoded in any of the three DNA molecules. However, because the poly-sugarphosphate backbone is always the same, it is doubtful that these DNA molecules convey genetic information.
- However, the nitrogen bases vary from one DNA segment to the next, therefore the information may depend on their sequences.
- In fact, the sequences of nitrogen bases in a specific section of DNA are similar to the linear sequence of amino acids in a protein molecule.
- An investigation of mutations of the head protein of bacteriophage T4 and the A protein of tryptophan synthetase from Escherichia coli provided the initial evidence for the colinearity between DNA nitrogen base sequence and amino acid sequence in protein molecules.
- Colinearity between protein molecules and DNA polynucleotides provides evidence that the arrangement of four nitrogen bases (e.g., A, T, C, and G) in DNA polynucleotide chains dictates the sequence of amino acids in protein molecules.
- These four DNA bases can therefore be viewed as the four alphabets of the DNA molecule. Therefore, all genetic information should be encoded using these four DNA alphabets.
- The question that now emerges is whether genetic information is written in articulated or coded language. If genetic information could have been communicated in an articulated language, the DNA molecule would have required multiple alphabets, a complicated grammar system, and adequate space.
- All of these could be practically difficult and also problematic for the DNA. Therefore, it was reasonable for molecular biologists to assume that genetic information resided in the DNA molecule as a specific language of code words that utilised the four nitrogen bases of DNA as their symbols. Any encoded message is referred to as a cryptogram.
Basis of Cryptoanalysis
- How information written in a four-letter language (four nucleotides or nitrogen bases of DNA) may be transformed into a twenty-letter language is the fundamental challenge of such a genetic code (twenty amino acids of proteins).
- A code word or codon is the set of nucleotides that specifies one amino acid. By genetic code, one refers to the collection of sequences of bases (codons) that correspond to each amino acid and translation signals.
- Regarding the possible size of a codon, we can consider George Gamov’s (1954) traditional yet rational explanation.
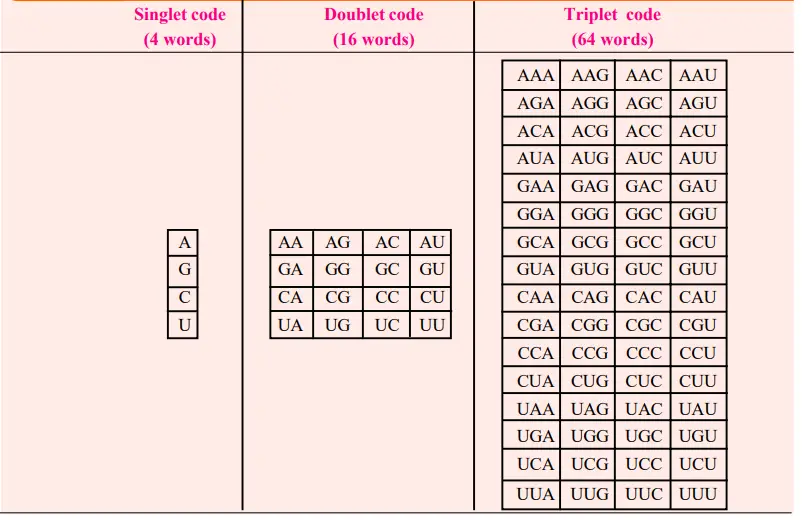
- The simplest conceivable code is a singlet code (a code of a single letter) that specifies a single nucleotide amino acid.
- A doublet code (consisting of two letters) is similarly insufficient, as it can only define sixteen (4×4) amino acids, but a triplet code (consisting of three letters) can specify sixty-four (4x4x4) amino acids.
- Therefore, it is probable that 64 triplet codes exist for 20 amino acids. The conceivable singlet, doublet, and triplet codes, which are conventionally described in terms of “mRNA language” [mRNA is a complementary molecule that copies the genetic information (cryptogram of DNA) during its transcription] are depicted in Table.
- In 1961, Crick and his colleagues present the first experimental evidence supporting the hypothesis of triplet coding.
- During their experiment, when they inserted or deleted single or double base pairs in a specific region of the DNA of E.coli T4 bacteriophages, they discovered that these bacteriophages ceased to execute their regular tasks.
- Nevertheless, bacteriophages with the addition or deletion of three base pairs in the DNA molecule had normal functionality.
- In this experiment, the addition of one or two nucleotides caused the message to be read incorrectly, however the addition of a third nucleotide resulted in the message being read correctly again.
Codon Assignment (Cracking the Code or Deciphering the Code)
The genetic code has been broken or deciphered using the following methods:
A. Theoretical Approach
- George Gamow, a physicist, proposed the diamond code (1954) and the triangle code (1955), as well as a comprehensive theoretical framework for the various aspects of the genetic code.
- Gamow proposed the following genetic code characteristics:
- A triplet codon that corresponds to a single polypeptide chain amino acid.
- Direct template translation by linking codons with amino acids.
- The code is translated in an overlapping fashion.
- Degeneration of the code, or the coding of an amino acid by more than one codon.
- The colinearity of nucleic acid and the produced main protein.
- Universality of the code, i.e., the code being fundamentally identical throughout organisms.
- Molecular biologists have refuted a number of these statements by Gamow. Brenner (1957) demonstrated that the overlapping triplet code is impossible, and further research has demonstrated that the code is non-overlapping.
- Crick’s adopter hypothesis similarly contested Gamow’s assumption of a direct template relationship between nucleic acid and polypeptide chain.
- Adaptor molecules, according to this concept, intervene between nucleic acid and amino acids during translation.
- In actuality, it is now understood that tRNA molecules serve as adaptors between the codons of mRNA and the amino acids of the resultant polypeptide chain.
B. The in vitro codon Assignment
1. Discovery and use of polynucleotide phosphorylase enzyme
Marianne Grunberg Manago and Severo Ochoa identified an enzyme from bacteria (e.g., Azobacter vinelandii or Micrococcus lysodeikticus) that catalyses RNA degradation in bacterial cells. The name of this enzyme is polynucleotide phosphorylase. Outside of the cell (in vitro), with high amounts of ribonucleotides, Manago and Ochoa discovered that the reaction could be driven in reverse and an RNA molecule could be produced (see Burns and Bottino, 1989). The random incorporation of nucleotides into the molecule is independent of a DNA template. Thus, in 1955, Manago and Ochoa made possible the artificial synthesis of polynucleotides (=mRNA) comprising only a single type of nucleotides (U, A, C, or G, respectively, repeated several times).
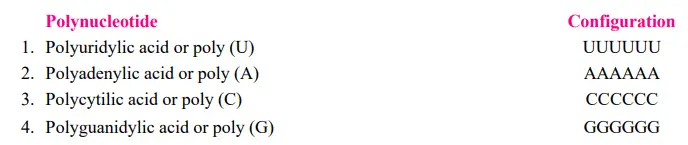
Consequently, the action of polynucleotide phosphorylase can be depicted as follows:
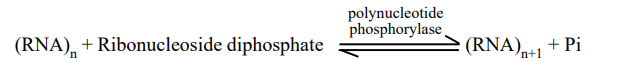
The polynucleotide phosphorylase enzyme differs from RNA polymerase used to transcribe mRNA and DNA polymerase used to transcribe mRNA from DNA in the following ways: I it does not require a template or primer; (ii) the activated substrates are ribonucleoside diphosphates (e.g., UDP, ADP, CDP, and GDP) and not triphosphates; and (iii (PPi). The introduction of synthetic (or artificial) polynucleotides and trinucleotides made the deciphering of the genetic code possible.
Use of polymers containing a single type of nucleotide (called homopolymers), mixed polymers (copolymers) containing multiple types of nucleotides (heteropolymers) in random or defined sequences, and trinucleotides (or “minimessengers”) in ribosome-binding or filter-binding are among the various techniques employed.
2. Codon assignment with unknown sequence
(i) Codon assignment by homopolymer.
- Marshall Nirenberg and Heinrich Matthaei (1961) supplied the first indication to codon assignment when they utilised an in vitro technique for the creation of a polypeptide utilising an artificially produced mRNA molecule containing only one type of nucleotide (i.e., homopolymer).
- Before doing the actual tests, they evaluated the capacity of a cell-free protein synthesis system to integrate radioactive amino acids into newly produced proteins.
- Their E.coli cell-free extracts comprised ribosomes, tRNAs, aminoacyl-tRNA synthetase enzymes, DNA, and messenger RNA.
- This extract’s DNA was eliminated by the deoxyribonuclease enzyme, so destroying the template for the synthesis of new mRNA.
- When twenty amino acids together with ATP, GTP, K+, and MG2+ were introduced to this mixture, they were integrated into proteins.
- As long as mRNA was present in the cell-free suspension, incorporation persisted. It also continued in the presence of synthetic polynucleotides (mRNAs) that might be synthesised using the polynucleotide phosphorylase enzyme.
- Nirenberg and Matthaei made the first successful application of this approach when they created a chain of uracil molecules (poly U) as their synthetic mRNA (homopolymer).
- A message consisting of a single base could not contain ambiguity, hence Poly (U) looked to be the best option. It binds well to ribosomes and, as it turned out, the resultant protein was insoluble and simple to isolate.
- When poly (U) was supplied as the message to the cell-free system containing all the amino acids, polyphenylalanine was picked solely from the mixture for incorporation into the polypeptide.
- This amino acid was phenylalanine, hence it was deduced that a sequence of UUU encoded for phenylalanine. Other homogeneous nucleotide chains (Poly A, Poly C, and Poly G) were inert for incorporation of phenylalanine. The phenlalanine mRNA code was consequently determined to be UUU.
- AAA is derived to be the equivalent DNA code word for phenylalanine. Thus, UUU was the first code word to be decrypted. In the laboratories of Nirenberg and Ochoa, this finding was developed.
- Using synthetic poly (A) and poly (C) chains, the experiment was repeated, yielding polylysine and polyproline, respectively.
- Thus, AAA was determined to be the code for lysine and CCC was determined to be the code for proline. A poly (G) message was discovered to be nonfunctional in vitro due to its secondary structure, which prevented it from attaching to ribosomes. Thus, three of the sixty-four codons were simply explained.
(ii) Codon assignment by heteropolymers (Copolymers with random sequences)
- Using synthetic messenger RNAs containing two different types of nucleotides, the genetic code was elucidated further.
- This approach was utilised in the laboratories of Ochoa and Nirenberg to deduce the codon composition for the 20 amino acids.
- The bases in the synthetic messengers were chosen at random (called random copolymers). In a random copolymer composed of U and A nucleotides, for instance, eight triplets are feasible, including UUU, UUA, UAA, UAU, AAA, AAU, AUU, and AUA.
- Theoretically, these eight codons may code for eight amino acids. However, actual experiments produced only six amino acids: phenylalanine, leucine, tyrosine, lysine, asparagine, and isoleucine.
- It was feasible to derive the composition of the code for different amino acids by altering the relative proportions of U and A in the random copolymer and determining the fraction of the different amino acids in the proteins generated.
3. Assignment of codons with known sequences.
- I The application of trinucleotides or minimessengers in filter binding (Ribosome-binding technique). Nirenberg and Leder’s (1964) ribosome binding technique takes use of the observation that aminoacyl-tRNA molecules attach selectively to the ribosomemRNA complex.
- The connection of a trinucleotide or minimessenger with the ribosome is necessary for aminoacyltRNA binding to occur.
- When a mixture of such small mRNA molecules-ribosomes and amino acid-tRNA complexes is incubated for a brief period and then filtered over a nitrocellulose membrane, the mRNA-ribosome-tRNA-amino acid complex is kept and the remainder of the mixture is discarded.
- Using a series of 20 different amino acid mixtures, each containing one radioactive amino acid, it is possible to determine the amino acid corresponding to each triplet by analysing the radioactivity absorbed by the membrane; for instance, the triplet GCC and GUU retain only alanyl-tRNA and valyl-tRNA, respectively.
- In this manner, all 64 potential triplets have been synthesised and evaluated. 45 of them have produced conclusive results. Later on, with the use of lengthier synthetic messages, 61 of the 64 potential codons have been deciphered.
C. The in vivo Codon Assignment
- Despite the fact that cell-free protein synthesis systems have played a significant role in the decipherment of the genetic code, they cannot tell us whether the deciphered genetic code is likewise utilised in the living systems of all organisms.
- Different molecular biologists use three techniques to determine if the same code is used in vivo: (a) amino acid replacement studies (e.g., tryptophan synthetase synthesis in E.coli and haemoglobin synthesis in man), (b) frameshift mutations (e.g., Terzaghi et al. 1966, on lysozyme enzyme of T4 bacteriophages), and (c) comparison of a DNA (e.g., comparison of amino acid sequence of the R17 bacteriophage coat protein with the nucleotide sequence of the R17 mRNA in the region of the molecule that dictates coat-protein synthesis by S. Cory et al., 1970).
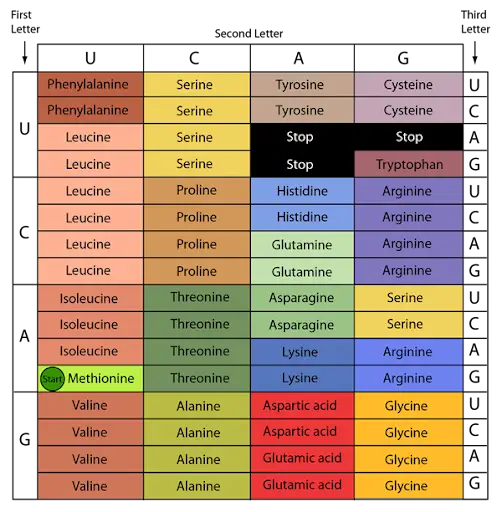
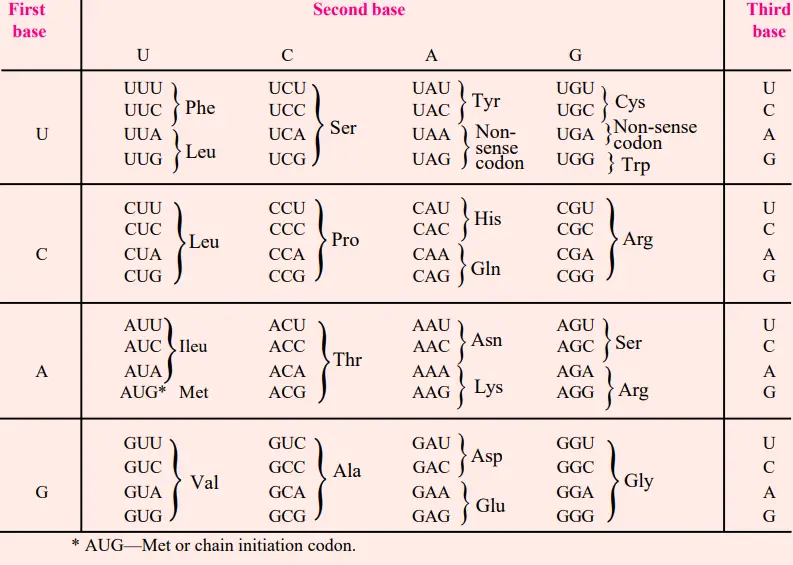
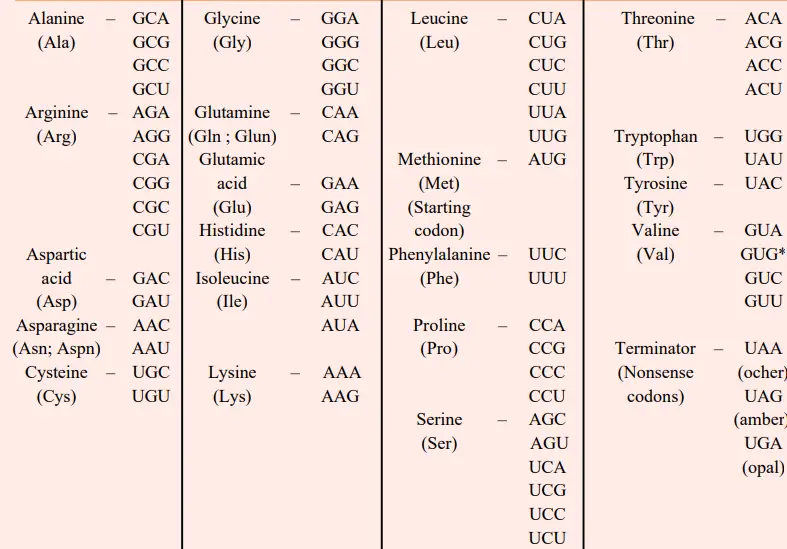
- Thus, the previously mentioned in vitro and in vivo experiments allowed for the formulation of a code table for twenty amino acids.
Characteristics of Genetic Code
The genetic code has the following general properties :
1. The code is a triplet codon
- The nucleotides of messenger RNA (mRNA) are organised as a linear sequence of codons, with each codon consisting of three consecutive nitrogenous bases, i.e., the code is a triplet codon.
- Two types of point mutations, frameshift mutations and base substitution, provide support for the concept of triplet codon.
(i) Frameshift mutations
- Evidently, the genetic communication, once launched at a particular place, is decoded into a series of three-letter phrases within a specific time frame.
- As soon as one or more bases are removed or added, the structure would be disrupted. When such frameshift mutations were intercrossed, they produced wild-type normal genes in certain combinations.
- It was determined that one was a deletion and the other was an insertion, so that the disordered frame order caused by the mutation will be corrected by the other.
(ii) Base substitution
- If, at a specific location in an mRNA molecule, one base pair is replaced by another without deletion or insertion, the meaning of a codon containing the altered base will be altered.
- As a result, another amino acid will be inserted in place of a particular amino acid at a particular location in a polypeptide.
- Due to a substitution mutation in the gene for the tryptophan synthetase enzyme in E. coli, the glycine-coding GGA codon becomes the arginine-coding AGA.
- A missense codon is a codon that has been altered to specify a different amino acid. The discovery that a fragment of mRNA comprising 90 nucleotides corresponded to a polypeptide chain having 30 amino acids of a developing haemoglobin molecule provided more direct proof for the existence of a triplet code.
- Similarly, 1200 nucleotides of the “satellite” tobacco necrosis virus direct the creation of 372 amino acid-containing coat protein molecules.
2. The code is non-overlapping
- In the translation of mRNA molecules, codons are “read” sequentially and do not overlap.
- Therefore, a non-overlapping coding indicates that a nucleotide in an mRNA is not utilised for multiple codons.
- In practise, however, six bases code for no more than two amino acids. In the event of an overlapping code, for instance, a single change (of replacement type) in the base sequence will result in several amino acid substitutions in the associated protein.
- In insulin, tryptophan synthetase, TMV coat protein, alkaline phosphatase, haemoglobin, etc., a single base substitution leads in a single amino acid change. Since 1956, a large number of examples have accumulated in which a single base substitution results in a single amino acid change.
- Recently, however, it has been demonstrated that overlapping genes and codons are possible in bacteriophage φ × 174.
3. The code is commaless
- The genetic code is punctuation-free, thus no codons are reserved for punctuation.
- It means that when one amino acid is coded, the next three characters will automatically code the second amino acid and no letters will be wasted as punctuation marks.
4. The code is non-ambiguous
- A codon always codes for the same amino acid when it is non-ambiguous.
- In the situation of ambiguous code, the same codon may have many meanings; in other words, the same codon may code for two or more amino acids. As a general rule, a single codon should never code for two distinct amino acids.
- There are, however, documented exceptions to this rule: the codons AUG and GUG may both code for methionine as beginning or starting codons, despite the fact that GUG is intended for valine. Similarly, the GGA codon represents the amino acids glycine and glutamic acid.
5. The code has polarity
The direction in which the code is always read is 5’→3′. Thus, the codon possesses a polarity. Clearly, if the code is read in opposing directions, it would specify two distinct proteins, as the codon’s base sequence would be reversed:
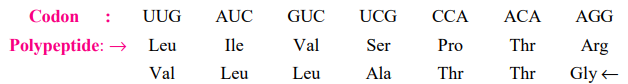
6. The code is degenerate
- Multiple codons might define the same amino acid; this phenomenon is known as degeneracy of the code. Except for tryptophan and methionine, which each contain a single codon, the remaining 18 amino acids have several codons.
- Consequently, each of the nine amino acids phenylalanine, tyrosine, histidine, glutamine, asparagine, lysine, aspartic acid, glutamic acid, and cysteine has two codons. Isoleucine consists of three codons.
- Each of the five amino acids valine, proline, threonine, alanine, and glycine has four codons. Each of the three amino acids leucine, arginine, and serine has six codons.
- There are essentially two types of code degeneration: partial and total. Partial degeneracy occurs when the first two nucleotides of degenerate codons are identical, but the third (3′ base) nucleotide differs, e.g., CUU and CUC code for leucine.
- Complete degeneracy happens when any of the four bases can code for the same amino acid in the third position (e.g., UCU,UCC, UCA and UCG code for serine).
- Degeneration of genetic coding has several biological benefits. It enables, for instance, bacteria with vastly different DNA base compositions to specify virtually the same complement of enzymes and other proteins.
- Degeneration also provides a technique for decreasing the lethality of mutations.
7. Some codes act as start codons
- In the majority of organisms, the AUG codon is the start or initiation codon, meaning that the polypeptide chain begins with methionine (eukaryotes) or N-formylmethionine (prokaryotes) (prokaryotes).
- Methionyl or N-formylmethionyl-tRNA binds particularly to the start site of mRNA with an AUG initiation codon.
- In rare instances, GUG functions as an initiating codon, such as in bacterial protein production. GUG normally codes for valine; however, when the regular AUG codon is deleted, only GUG is used as an initiation codon.
8. Some codes act as stop codons
- Triple codons UAG, UAA, and UGA are the stop or termination codons for the chain. They do not code for any of the amino acids.
- These codons are not read by any tRNA molecules (via their anticodons), but are read by some specialised proteins, called release factors (e.g., RF-1, RF-2, RF-3 in prokaryotes and RF in eukaryotes) (e.g., RF-1, RF-2, RF-3 in prokaryotes and RF in eukaryotes).
- These codons are also called nonsense codons, since they do not designate any amino acid. The UAG was the first termination codon to be found by Sidney Brenner (1965). (1965).
- It was named amber in honour of a doctoral student named Bernstein (= the German term for ‘amber,’ and amber signifies brownish yellow) who helped identify a class of mutations.
- Apparently, the other two termination codons were also named after colours, such as ochre for UAA and opal or umber for UGA, in order to maintain consistency. (ochre indicates pale yellow or golden red, opal means milky white, and umber signifies brown)
- The presence of multiple stop codons may be a precautionary mechanism in case the first stop codon fails to work.
9. The code is universal
- The same genetic code is valid for all creatures, from bacteria to humans. Marshall, Caskey, and Nirenberg (1967) showed the universality of the code by showing that E.coli (bacterial), Xenopus laevis (amphibian), and guinea pig (mammal) amino acyl-tRNA utilise nearly the same code.
- Nirenberg has also suggested that the genetic code may have originated with the first bacteria three billion years ago, and that it has altered very little over the history of living species.
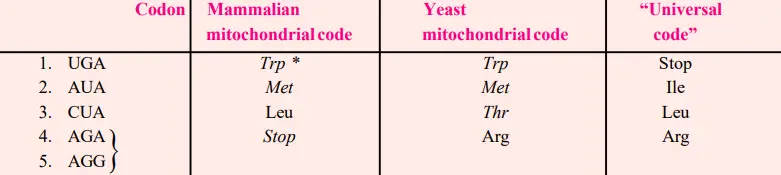
- Recently, inconsistencies between the universal genetic code and the mitochondrial genetic code have been revealed.
Codon and Anticodon
- The codon words of DNA are complementary to the mRNA code words (i.e., DNA codes run in the 3’→5′ direction whereas mRNA code words run in the 5’→3′ direction), as are the three bases composing the anticodon of tRNA (i.e., anticodon bases run in the 3’→5′ direction).
- Three bases of the anticodon pair with the mRNA on the ribosomes during the alignment of amino acids during protein synthesis (i.e., the translation of mRNA into proteins in the N2→COOH direction).
- For instance, one of the two mRNA and DNA code words for the amino acid phenylalanine is UUC, while the equivalent anticodon of tRNA is CAA.
- This suggests that the pairing of codons and anticodons is antiparallel. C pairs with G and U pairs with A in this instance.
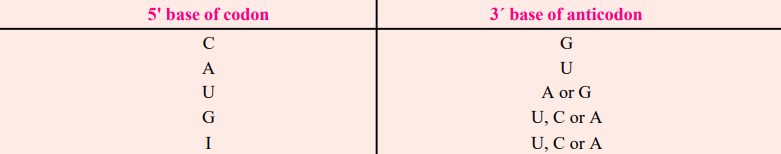
Wobble Hypothesis
- Crick (1966) presented the wobble hypothesis to explain the potential origin of codon degeneracy (wobble means to sway or move unsteadily).
- Given that there are 61 codons that specify amino acids, the cell must possess 61 tRNA molecules, each with a unique anticodon.
- The actual number of tRNA molecule types discovered is far fewer than 61. This suggests that tRNA anticodons read many codons on mRNA.
- For instance, yeast tRNAala with anticodon bases 5′ IGC 3′ (where I stands for inosine, a derivative of adenine or A) may bind to three codons in mRNA, including 5′ GCU 3′, 5’GCC3′, and 5′ GCA3′.
- Inosine is usually found as the 5′ base of the anticodon; when pairing with the base of the codons, it wobbles and can pair with U, C, or A of three different codons.
- Therefore, according to Crick’s wobble hypothesis, the base at the 5′ end of the anticodon is not as spatially constrained as the other two bases, allowing it to establish hydrogen bonds with any of the bases positioned at the 3′ end of a codon.