Table of Contents
- Following the work of Meselson and Stahl, researchers confirmed that other organisms use semiconservative replication. However, semiconservative replication can occur in a number of distinct ways, distinguished primarily by the nature of the template DNA (linear or circular) and the number of replication forks.
- Replicons are the individual units of replication, and each one contains a replication origin. Beginning at the origin, replication proceeds until the complete replicon has been reproduced. Eukaryotic chromosomes include several replication origins, whereas bacterial chromosomes contain only one.
What is Theta Model of Replication?
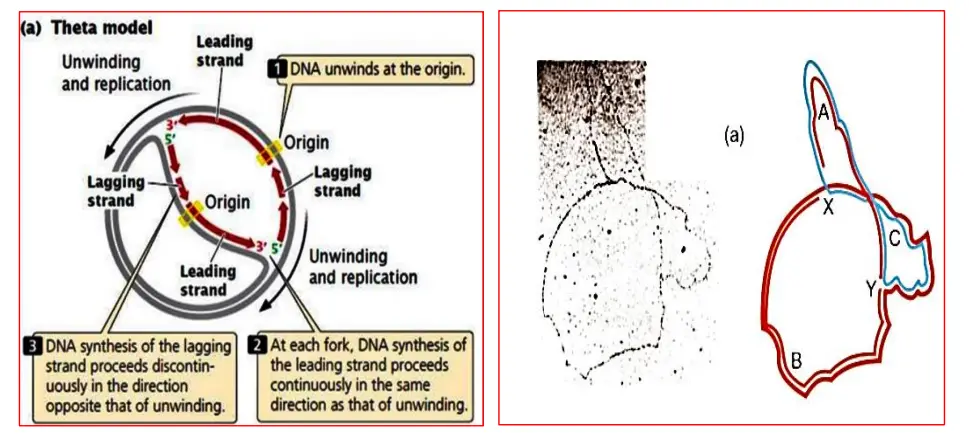
Theta structures are produced during the replication of circular DNA molecules (prokaryote DNA). Two replication forks can move independently around the DNA ring, and the structure resembles the Greek letter (theta) when viewed from above.
- DNA molecules that are circular exhibit the theta mode of replication.
- Circular chromosomes are a type of circular DNA found in bacteria and archaea that lacks free ends, in contrast to the linear DNA strands present in the majority of eukaryotes.
- In circular DNA, there are three types of replication modes: rolling-circle, strand displacement, and theta.
- Similar to the linear form of chromosome replication, the theta mode of replication replicates leading strands constantly and trailing strands intermittently.
- No DNA breaks are necessary for this method of replication.
- The theta method of replication is prevalent in bacteria, such as Escherichia coli and Bacillus subtilis, chloroplasts, and mitochondria.
- Because of an intermediate form that resembles the Greek letter θ, this style of replication is known as the theta mode.
- It was discovered by John Cairns and led to the realisation that DNA replication may occur in both directions.
Procedure of Theta (θ) Model of Replication
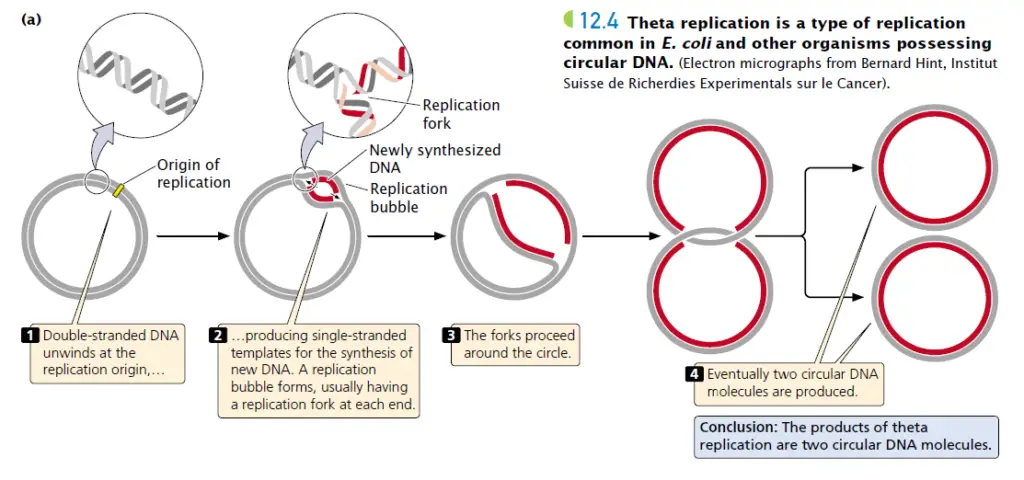
There are three phases to the replication of circular DNA: initiation, elongation, and termination.
1. Initiation
- The replication process begins at a conserved sequence region referred to as the origin of replication or oriC.
- At the oriC, the assembly of initiator proteins begins.
- The regulation of the assembly of initiator proteins ensures that replication occurs only once every cell cycle.
2. Elongation
- During elongation, the proteins at oriC begin to travel in the opposite direction around each arm of the chromosome.
- This mechanism, known as bidirectional replication, generates two identical copies of the chromosome.
- Each arm of the chromosome is called a replichore, and the entire protein-containing structure is called a replisome.
- In the front of the replisome is the enzyme DNA helicase, which unwinds the two strands of circular DNA and generates a replication fork.
- The unwound strands are then used as a template by DNA polymerase and other assembling proteins to synthesise the complementary strand.
- When the replication fork moves around the circular chromosome during replication, a theta-shaped structure is created.
- John Cairns, a British physician, demonstrated this by radioactively labelling E.coli chromosomes with 3H-thymidine and observing them under X-ray on an electron micrograph grid.
3. Termination
- On circular chromosomes, opposite the oriC is the terminus region, which consists of several “Ter” sites.
- The replication terminator protein halts the two replication forks at the Ter site and brings them to a single chromosomal location.
- Daughter chromosomes are segregated while the enzyme machinery is dismantled.
What is Cairn’s experiment?
- In 1968, J. Cairns used a combination of microscopy and autoradiography to make a method that made it possible to see the E. coli chromosome copying itself.
- Cairns put E. coli cells in a medium with 3H-thymine in it (tritiated -thymidine was incorporated into the DNA as the chromosome was replicated in successive generations of cells).
- After different times of incubation, the E. coli were taken out of the medium and gently lysed to get the chromosome out of the cell (since the shear forces created by harsh lysis break the chromosome into small pieces).
- The chromosomes were then put on glass slides and covered with a photographic emulsion that was sensitive to the 3Hthymidine’s low-energy beta particles.
- After the beta rays were shone on the emulsion, it was developed and looked at with a light microscope.
- Wherever the labelled thymidine in a chromosome had broken down, the emulsion was exposed and made visible grains.
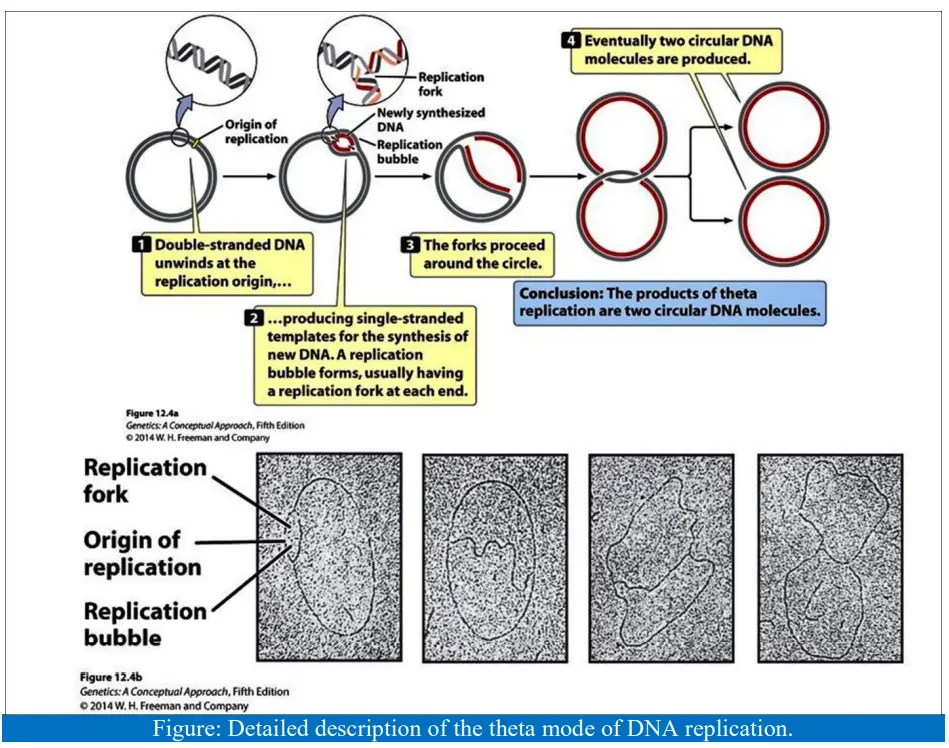
- A chromosome that wasn’t replicating looked like a circle made of exposure spots that were close together.
- Theta configurations were made when chromosomes were “caught in the act” of copying themselves. They look like the Greek letter theta.
- In circular chromosomes, the theta structures show where the replication forks are.
- The results showed that the radioactive thymidine was added to both forks of the theta structure, not just one. This shows that synthesis is happening at both forks in opposite directions around the loop. Because of this, replication can go both ways.
From John Cairns Experiment on E. coli it was found that-
- In this process of replication, the circular DNA keeps its shape. During this process, the circle doesn’t seem to be broken.
- Because of the replication eye, a Theta structure in the middle is made.
- The replication seems to happen at one or two moving Y-junctions in the circle replication forks, making it a semiconservative type.
References
- Lilly J, Camps M. Mechanisms of Theta Plasmid Replication. Microbiol Spectr. 2015 Feb;3(1):PLAS-0029-2014. PMID: 26005599; PMCID: PMC4441207.
- https://onlinelibrary.wiley.com/doi/abs/10.1128/9781555818982.ch3
- https://en.wikipedia.org/wiki/Theta_structure
- https://sacbiotech.files.wordpress.com/2017/11/the-theta-mode-of-dna-replication-in-escherichia-coli.pdf
- https://www.ritubiology.com/2016/07/19/dna-replication-what-is-dna-replication-theta-model-rolling-circle-replication/
- https://plantlet.org/replication-in-circular-dna-theta-model/
- https://byjus.com/neet/theta-mode-of-replication/#:~:text=The%20theta%20mode%20of%20replication,strands%20seen%20in%20most%20eukaryotes.
