Table of Contents
- Clostridium perfringens is a gram-positive, rod-shaped bacterium that is responsible for causing frequent foodborne diarrheal disease. It is a commonly occurring foodborne illness, particularly in industrialized nations.
- Clostridium perfringens can be found ubiquitously in the environment and is pathogenic to both humans and animals. One of the severe diseases caused by this bacterium is human enteritis necroticans, also known as pigbel, which can be life-threatening and leads to ulceration of the small bowel.
- As a spore-forming anaerobic bacterium, Clostridium perfringens produces a toxin called C. perfringens enterotoxin (CPE) during sporulation. Different strains of Clostridium perfringens, classified as Types A to E, are associated with different diseases.
- The biological properties of CPE-producing strains include their ability to rapidly multiply within a short period, grow at a relatively high range of temperatures, form spores that can withstand harsh environmental conditions, and tolerate the presence of oxygen despite being anaerobes.
- It is important to note that there are other pathogenic species within the Clostridium genus, such as Clostridium tetani (causing tetanus), Clostridium botulinum (causing botulism), Clostridium baratii, Clostridium argentinese, and Clostridium difficile (associated with antibiotic-associated diarrhea and pseudomembranous colitis).
Characteristics of Clostridium perfringens
- Clostridium perfringens is a Gram-positive, nonmotile, rod-shaped bacterium that is responsible for causing various human and veterinary illnesses.
- The pathogenicity of Clostridium perfringens is primarily attributed to its ability to produce toxins. More than 16 distinct toxins have been identified in isolates associated with human and veterinary illnesses.
- In human gastrointestinal (GI) diseases caused by Clostridium perfringens, three toxins play a significant role: CPE (C. perfringens enterotoxin), beta-toxin (the main toxin responsible for necrotizing enteritis in humans), and possibly beta2-toxin (an additional toxin implicated in human colitis caused by CPE-producing type A strains).
- While toxin production is crucial in human GI diseases, Clostridium perfringens isolates associated with food poisoning possess additional biological characteristics. These include a rapid doubling time of ten minutes for vegetative cells, the ability of vegetative cells to grow at relatively high optimal temperatures (43-45°C), the formation of stress-resistant spores, and the ability to tolerate exposure to air despite being classified as an anaerobic bacterium.
- These features contribute to the development of Clostridium perfringens foodborne diseases. Clostridium perfringens isolates are classified into five types (A-E) based on the synthesis of four main toxins (alpha, beta, epsilon, and iota). Traditionally, toxin-antitoxin serum neutralization experiments in mice have been used for strain typing, but PCR-based genotyping systems have simplified the process.
- Type A strains are the most common in food poisoning cases, while the cpe gene, which encodes CPE, is infrequently found in type C, D, and E strains. Although type C to E strains may produce an enterotoxin similar to CPE, they have not been definitively identified as causative agents of C. perfringens food poisoning outbreaks.
- CPE-producing type A strains carrying the cpe gene are more widespread than other CPE-producing strains among the different types (A-E). However, there have been isolated cases of necrotic enteritis (pigbel) caused by cpe-positive type C strains, and accidental isolation of cpe-positive type C strains from patients with type A food poisoning has also been reported.
- In most, if not all, type C, D, and E strains, the cpe gene is carried on a conjugative transferrable plasmid. The genetic arrangement of the cpe region in these strains differs from that of cpe-positive type A strains.
Factors Contributing to C. perfringens Type A Foodborne Disease
- C. perfringens type A foodborne disease is primarily attributed to the presence of CPE (C. perfringens enterotoxin), a unique toxin that plays a significant role in the gastrointestinal pathogenicity of the bacterium.
- Numerous epidemiological investigations have demonstrated the association between CPE and food poisoning outbreaks caused by type A C. perfringens isolates. The presence of CPE is strongly correlated with the occurrence of the disease, and experimental animal studies have shown that the level of CPE in a patient’s feces can induce severe intestinal consequences.
- Human volunteers who have been administered highly purified CPE have experienced all the symptoms associated with C. perfringens type A food poisoning.
- Animal model research has shown that CPE acts on the gastrointestinal tract, affecting all sections of the small intestine. Purified CPE leads to rapid fluid and electrolyte loss from the GI tract in rabbit ileal loops. This effect can be counteracted by CPE-specific antiserums.
- Furthermore, CPE-positive C. perfringens food poisoning isolates induce greater fluid buildup and histopathologic damage in rabbit ileal loops compared to CPE-negative isolates or a cpe knockout mutant of the food poisoning strain.
- The fact that CPE-producing type A C. perfringens strains, most of which carry the cpe gene on a plasmid, are also found in non-foodborne human GI diseases such as antibiotic-associated diarrhea, sporadic diarrhea, and nosocomial diarrheal outbreaks further supports the significance of CPE in human GI virulence.
Source of contamination of Clostridium perfringens
- C. perfringens can be found in various environmental sources such as soil, water, sludge, sewage, and contaminated equipment. However, it is primarily associated with foodborne contamination.
- Raw and frozen meats, particularly pork, beef, and poultry products cooked with sauces, pose a high risk of C. perfringens contamination. These types of foods provide a favorable environment for the bacterium to grow and multiply.
- Certain settings such as school canteens, hospitals, restaurants, prisons, and gatherings where large quantities of food are prepared and served are at a higher risk of Clostridium outbreaks.
- Additionally, C. perfringens can be found in the intestinal tracts of humans and animals, particularly in the feces of the elderly. Poor personal hygiene and inadequate sanitation practices can contribute to the spread of the bacterium from fecal sources to food.
- C. perfringens requires various amino acids and vitamins for growth and thrives at temperatures between 43-45°C. Protein-rich foods, such as meat, serve as an excellent reservoir for its growth. If food is not adequately cooled or allowed to remain at room temperature during the heating and cooling process, C. perfringens can proliferate and produce toxins, increasing the risk of contamination and subsequent food poisoning.
Clostridium perfringens enterotoxins
- Clostridium perfringens produces enterotoxins during sporulation, typically under stressed environmental conditions. Among the various toxins it produces, there are five enterotoxins designated as Type A to E.
- C. perfringens also produces four types of extracellular toxins: alpha (α), beta (β), epsilon (ε), and iota (Ϊ).
- Type A enterotoxin, known as CPE (Clostridium perfringens enterotoxin), is associated with several diseases in humans and animals. In humans, it can cause gas gangrene and antibiotic-associated infectious diarrhea. In animals, Type A CPE is linked to enterotoxemia, necrotic enteritis, and acute gastric dilation.
- Type B, D, and E enterotoxins have not been definitively associated with human diseases, but they can cause enteritis and enterotoxemia in animals.
- Type C enterotoxin, similar to Type A, can lead to necrotizing enteritis (jejunitis) in humans. It also causes enterotoxemia and necrotic enteritis in piglets, lambs, calves, and birds.
- In addition to enterotoxins, C. perfringens produces other toxins such as perfringolysin O (theta toxin) and collagenase (kappa toxin).
- These various toxins play a role in the pathogenicity of C. perfringens, contributing to the symptoms and severity of the associated diseases in both humans and animals.
Epidemiology of Clostridium perfringens food poisoning
- Clostridium perfringens enterotoxins are associated with foodborne outbreaks that have been reported in various regions including the United States, Europe, and Japan. These outbreaks typically involve an average of around 50 to 100 individuals.
- Several factors contribute to the occurrence of C. perfringens outbreaks. These include the preparation of food in large quantities, inadequate cooking temperatures (specifically, insufficient cooking of the core region of the food), and contamination during the period between food preparation and serving.
- Outbreaks frequently occur in settings where a significant amount of food is prepared in advance, such as hospitals, school canteens, prisons, parties, seminars, and gatherings.
- For example, there was an outbreak in a mental hospital that affected 170 patients and resulted in the death of 3 patients. The investigation revealed that the source of the outbreak was a lamb dish that had been cooked and stored at room temperature for 20 hours before being served without reheating the meat. C. perfringens was isolated from the lamb meat, infected fecal samples, and post-mortem samples.
- Another outbreak occurred in a school canteen due to the consumption of a C. perfringens-contaminated milkshake. The milkshake had been left at room temperature after preparation, leading to 77 cases of diarrhea.
- These outbreaks highlight the importance of proper food handling and storage practices, particularly in terms of temperature control, to prevent the growth and toxin production of C. perfringens and subsequent foodborne illnesses.
Synthesis and Release of CPE
- Although C. perfringens isolates are classified into three cpe genotypes, the amount of CPE generated by an isolate is unaffected by whether the cpe gene is located on the chromosome or on a big plasmid.
- CPE synthesis begins shortly after sporulation stimulation and gradually increases over the next 6-8 hours.
- CPE expression is transcriptionally controlled, with considerable amounts of cpe mRNA produced during sporulation; however, cpe mRNA is not produced during C. perfringens vegetative growth.
- This phenomena is highly supported by the following genetic analysis findings: In type A strains, sporulation-associated alternative sigma factors such as SigF, SigK, and SigE tightly regulate cpe gene expression. SigK and SigE (sporulation-associated sigma factors active in mother cells during Bacillus subtilis sporulation) directly activate cpe mRNA transcription by engaging SigK- or SigE-dependent promoters (P1, P2, and P3) positioned upstream of the cpe ORF.
- The high level of CPE expression during sporulation is also due to the extraordinary stability of cpe mRNA (the functional half-life of cpe mRNA is approximately 60 min).
- CPE, unlike most C. perfringens toxins, is not actively released; instead, it accumulates in the cytoplasm of the mother cell as a cytoplasmic CPEcontaining paracrystalline inclusion body after synthesis.
- When the sporulation activities are complete, the mother cell lyses, releasing CPE into the intestinal lumen, where it acts as an enterotoxin.
Effects of CPE on the GI Tract
- In humans, C. perfringens food poisoning is a self-limiting sickness. As a result, histopathologic research on humans have been rare.
- Rather, the effects of CPE have been studied using animal models. CPE causes fluid and electrolyte loss from the GI tract and then causes extensive histopathologic damage to the small intestine in an in vivo rat or rabbit ileal loop model; thus, the target organ for CPE is thought to be the small intestine, with the ileum being particularly sensitive to this toxin.
- CPE-induced histopathologic damage is initially limited to the villus tips of the small intestine; however, over time, the entire small intestinal villus suffers extensive damage, including desquamation of the villus epithelium, inflammatory cell infiltration (primarily lymphocytes), and mild hyperemia in the mucosa.
- The initiation of fluid transport modifications closely coincides with the development of tissue damage, and CPE levels that cause tissue damage can induce intestinal fluid and electrolyte transport variations.
- CPE-induced intestinal damage can occur in the rabbit ileum within 15-30 minutes of toxin treatment. CPE’s biological activities were studied in depth utilising a cultivated intestinal epithelial cell line.
- These studies hypothesised that CPE would operate in a multistep mechanism. The current paradigm for CPE action begins with temperature-sensitive binding to the toxin-specific receptors claudin-3 and claudin-4.
- Claudins are a 24-member protein family ranging in size from 20 to 25 kDa that are the most significant proteins in epithelial tight junctions.
- These tight junction proteins are projected to have four transmembrane domains, two extracellular loops, and signal cascades mediated by the cytoplasmic tail.
- CPE binds to receptor claudin proteins via the tight junction protein’s second extracellular loop (at a helix-turn-helix motif). CPE can bind to claudins-3, claudin-4, claudin-6, claudin-7, claudin-8, and claudin-14 (claudins-8 and claudin-14 contribute less to CPE cytotoxicity than claudin-4), but not to claudins-1, claudin-2, claudin-5, or claudin-10.
- CPE-claudin binding causes the creation of a 90 kDa small complex containing claudin families and claudin receptors, as well as claudins incapable of binding CPE (nonreceptor claudin(s)).
- Six tiny complexes are thought to oligomerize into a stable big complex (450 kDa) after 5 minutes of CPE binding, which first assembles as a prepore and rapidly inserts into membranes to create an active pore.
- The creation of active holes results in a lack of normal plasma membrane permeability. These CPE-induced permeability changes are initially limited to tiny molecules with 200 Da, such as calcium ions and amino acids.
- CPE-treated cells die via traditional caspase 3-mediated apoptosis at low levels of CPE, whereas cells die from oncosis at greater levels of CPE as a result of these processes.
- Thus, large-complex formation is required for CPE-induced cytotoxicity. CPE pore development also causes morphological damage, allowing for the establishment of a larger large complex (650 kDa).
- Occludin, a 65 kDa tight junction protein, is included in this 650 kDa big complex. This incident causes tight intersections to be further disrupted.
- The involvement of tight junction proteins in CPE activities results in alterations in intestinal paracellular permeability.
- The complete sequence of events in CPE-treated culture cells is thought to result in CPE’s effects on the small intestine in vivo and in vitro.
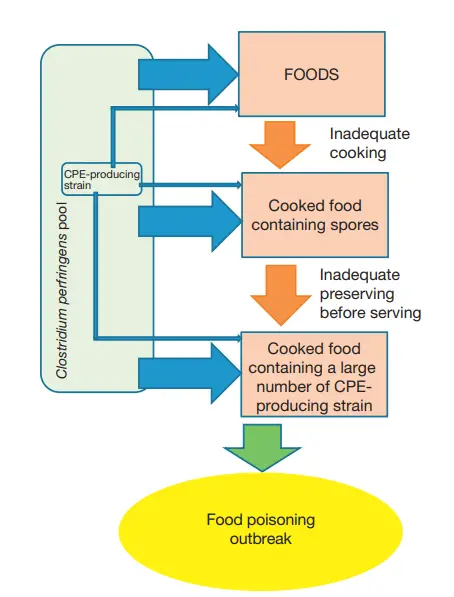
Pathogenesis of C. perfringens Food Poisoning
- Inappropriate heating and preservation cause the vegetative cells of enterotoxigenic C. perfringens in foods to proliferate rapidly.
- While the acidity of the stomach certainly kills many eaten C. perfringens vegetative cells, some vegetative cells that survive pass past the stomach and remain viable in a sufficiently contaminated food carrier (i.e., food containing 106 –107 C. perfringens vegetative cells per gram).
- Because the surviving vegetative cells reproduce and sporulate in the small intestine, this condition is classified as an infection rather than an intoxication.
- A sporulation factor with a low molecular weight (1000-5000) generated by both CPE-negative and CPE-positive vegetative cells may induce in vivo sporulation.
- CPE is expressed during sporulation and accumulates in C. perfringens mother cells.
- CPE is released into the intestinal lumen when mother cells lyse, where it immediately attaches to intestinal epithelial cells and exerts its function.
- These occurrences result in fluid and electrolyte losses, as well as morphological damage to intestinal epithelial cells.
Source of Enterotoxigenic C. perfringens Strains in Food
- While meat and meat products (particularly beef and poultry and gravies) were initially thought to be common food vehicles for C. perfringens type A foodborne illness in the United States, various foods have also been identified as a contaminated food in C. perfringens type A food poisoning outbreaks.
- C. perfringens is found in soil (at levels of 103 -104 CFU g-1), foods (e.g., approximately 50% of raw or frozen meat has some amount of C. perfringens), dust, and the intestinal tracts of humans and domestic animals (e.g., human faeces typically contain 104 -106 CFU g-1).
- Independent surveys studying diverse types of environmental samples found both types (chromosomal or plasmid-borne) of cpe-positive isolates, whereas other surveys failed to find cpe-positive bacteria.
- The following are the reasons for the difficulties in finding C. perfringens generating CPE in environmental samples:
- C. perfringens is found in very low numbers in retail food samples, whereas the C. perfringens strain is found often in retail food.
- Only a small percentage of dietary isolates (less than 5%) can generate CPE, possibly even in putative reservoirs (s).
- CPE is only formed during sporulation; however, it can be challenging to induce sporulation of isolates using various sporulation-specific media (e.g., Duncan–Strong medium).
- The small number of surveys identifying cpepositive strains in studied samples suggested that C. perfringens strains harbouring the cpe gene on a large plasmid are likely to be the major population in the environment, whereas the three C. perfringens cpe genotypes are widely spread.
- As a result, cpe-positive bacteria from the environment could be accidently contaminated in any step of food production.
- Recent molecular assay studies (multilocus sequence typing and microarray assays) have revealed the genetic features of chromosomal and plasmid cpe strains.
- The DNA sequences of roughly 450-500 bp internal segments of various housekeeping genes were used in the multilocus sequence typing (MLST) technique to classify isolates.
- The various sequences present within isolates of each housekeeping gene have been designated as separate alleles, and the alleles at each of the loci define the allelic profile or sequence type for each isolate (ST).
- MLST assays are used to distinguish chromosomal cpe strains from plasmid cpe strains, cpe-negative strains, and type B to E veterinary strains.
- A microarray assay is another molecular assay in which nucleic acid sequences specifically pair with each other between complementary nucleotide base pairs.
- Type A chromosomal cpe strains have different genetic backgrounds than type A plasmid cpe strains and cpe-negative type A strains, according to a completely sequenced C. perfringens food poisoning strain genome-based DNA microarray assay; that is, the gene clusters of myo-inositol, ethanolamine, and biotin synthesis are absent in the variable region of chromosomal cpe strains, but
- When the results of MLST analysis for chromosomal cpe strains are combined, chromosomal cpe strains and plasmid cpe strains are likely to have different habitats in the environment; plasmid cpe strains may have adapted to the mammalian intestine environment, whereas chromosomal cpe strains may be present in environments with plant materials.
- The primary reservoir(s) of C. perfringens type A food poisoning strains have not been thoroughly investigated. To further investigate the source of CPE-producing C. perfringens in environmental samples, a method(s) capable of detecting low numbers of bacterial cells is required, because contaminated samples commonly contain a low number of CPE-producing C. perfringens with a small subset of total C. perfringens isolates.
- Unfortunately, the current conventional techniques for diagnosing C. perfringens food poisoning can only detect a small fraction of cpe-positive strains in samples. Several tests based on’microbial source tracking’ approaches could be beneficial for detecting low numbers of vegetative cells and/or spores in environmental samples.
- In the future, epidemiological surveys using assays that can distinguish between the genetic backgrounds of cpepositive and cpe-negative strains and are based on microbial source tracking will aid in understanding the cpe-positive C. perfringens habitat in the environment, and candidate(s) of native reservoirs of chromosomal and/or plasmid cpe-positive C. perfringens strains will be identified based on these epidemiological findings.
Clinical Features of Clostridium perfringens Food Poisoning
- Food poisoning outbreaks caused by type A enterotoxigenic C. perfringens are often considerable in number. Food poisoning is characterised by fairly severe stomach cramps, nausea, and watery diarrhoea; vomiting and fever are uncommon.
- Food poisoning symptoms caused by type A C. perfringens strains appear 8-24 hours after consuming substantially contaminated food.
- The disease is usually self-limiting, lasting 12-24 hours.
- Mild symptoms can continue for a week or two in certain circumstances. Fatalities are extremely rare, occurring in less than 0.05% of instances.
- While everyone is susceptible to C. perfringens type A food poisoning, dehydration is the leading cause of death, which occurs in the very young, very elderly, disabled, and chronically unwell.
Detection/Diagnosis of Clostridium perfringens
The determination of CPE in patient faeces and bacteriological examinations of stools and incriminated food are used to identify C. perfringens food poisoning. The following are the bacteriological criteria:
- Food containing a high concentration of vegetative C. perfringens cells (>105 per gramme).
- The organism was isolated in great quantities (>106 per gramme) from faecal specimens. The normal human population has a faecal count of 103 per gramme. Several reports, however, indicate that C. perfringens spore counts greater than 106 per gramme can also be discovered in debilitated, institutionalised patients who are neither severely unwell or implicated in a food-poisoning outbreak.
Additional investigations to link human sickness with tainted food include the following:
- The identification of a common C. perfringens toxinotype using polymerase chain reaction (PCR) detection of all toxin genes, pulse-field gel electrophoresis (PFGE), multilocus sequence typing (MLST), or ribotype in faecal specimens and the incriminated food.
- A toxinotype, PFGE, MLST, or ribotype found in faecal collections from multiple people.
Clostridium perfringens Enterotoxin Assays
- Because CPE is only formed during sporulation, CPE detection in culture supernatant requires cultivation in a particular sporulation medium and control of the presence of sporulating cells.
- Several sporulation medium have been proposed, with varying success depending on the strain.
- A typical C. perfringens sporulation protocol is as follows: A 1 ml culture of C. perfringens grown in cooked beef medium is transferred to a 10 ml fluid thioglycollate medium.
- Heat shock the inoculated fluid thioglycollate media for 20 minutes at 70 C. The fluid thioglycollate culture is transferred to 100 ml Duncan-Strong sporulation medium and incubated at 37 degrees Celsius overnight.
- The culture is examined under phase-contrast microscopy for the presence of spores, and the culture supernatant collected by centrifugation is subjected to CPE detection.
- CPE can also be identified directly in faecal samples prepared as follows. In a vortex mixer, one volume (1 ml) of faecal material (about 1 g) is combined with one volume (1 ml) of 0.001 M phosphate buffer pH 7.2, containing 0.15 M sodium chloride (phosphate buffered saline (PBS)).
- The suspension is centrifuged at 12 000 g for 20 minutes at 4 C, or it is passed through 0.45 or 0.22 mm membrane filters, and the supernatant or filtrate is examined. Initially, biological approaches such as mouse lethality, Vero cell cytotoxicity, and Vero cell plating inhibition were utilised to identify CPE.
- Specific polyclonal and monoclonal anti-CPE antibodies have been produced, and a wide range of immunological tests for CPE detection and titration have been proposed.
- The initial immunological tests were based on CPE immunoprecipitation on agarose gel in the presence of particular antibodies: single-gel diffusion and double-gel diffusion, also known as the Ouchterlony test.
Counterimmunoelectrophoresis
- The introduction of an electrical field improves the sensitivity of precipitation processes (Table 3), and counterimmunoelectrophoresis is the most often utilised of these approaches.
- In agarose gel, two rows of wells separated by roughly 5 mm are cut.
- Serial dilutions of CPE and samples are spread in one row’s wells, while anti-CPE antibodies are scattered in the other row’s wells.
- For 30-60 minutes, an electrical field (10 V cm1) is applied (near the antigen-containing wells).
- In the presence of CPE, a precipitation channel is visible.
Latex Agglutination Tests
- There are two latex agglutination tests: reverse passive latex agglutination (RPLA), which requires overnight incubation, and slide latex agglutination (SLAT), which takes only a few minutes.
Reverse Passive Latex Agglutination
RPLA is a commercially available product (PET-RPLA, TD930, Oxoid, Basingstoke, UK). The sensitivity is around 3 ng ml1. The steps are as follows:
- Two rows of a 96-well V type microtiter plate are utilised for each sample.
- Except for the first well of each row, fill each well with 25 ml of PBS containing 9.5% bovine serum albumin (BSA). The final wells contain solely PBS-BSA.
- Fill the first and second wells of each row with a 25 ml sample.
- From the second to the seventh well, serial twofold dilutions are performed.
- To the first row of wells, add 25 ml of beads sensitised with immunopurified antiCPE antibodies.
- In each well of the second row, add 25 ml of control beads sensitised with nonimmune rabbit immunoglobulins.
- Mix thoroughly by hand rotating the plate or using a plate shaker.
- Place the microplate in a humidified chamber or cover it with a lid.
- Incubate the plate for 20-24 hours at room temperature.
The following is how the results are interpreted:
- Visual inspection is used to determine agglutination. This is made easy by placing a black sheet beneath the microplate or using a test reading mirror.
- The scores are +++ (full agglutination), ++, +, +/, or -. (absence of agglutination).
- Control latex must be present in a negative row. In some instances, nonspecific agglutination might be seen. When the positive agglutination in the sensitised row surpasses that in the control row by two wells or more, the sample is considered to have CPE.
Slide Latex Agglutination
The SLAT technique involves agglutinating latex beads on a glass slide in the presence of CPE. The following steps are taken to prepare the reagent:
- 1:3 dilution of latex beads (0.8 mm) in glycine buffer (0.1 M glycine, 0.15 M NaCl, pH 8.2).
- Add anti-CPE immunoglobulins isolated by immunoaffinity on a Sepharose column with immobilised CPE (13 mg ml1 , final concentration).
- After 1 minute of room temperature agitation, add an equal amount of PBS-0.1% BSA and vortex vigorously to combine the suspension.
- For the control latex, use nonimmune rabbit immunoglobulin G (Sigma). 5. Keep the latex suspensions at 4 degrees Celsius.
The following is the test procedure:
- On a glass slide, combine 25 ml of samples and repeated twofold dilutions in PBS containing 0.1% BSA with 25 ml of sensitised or control latex beads. After 1-5 minutes, gently rotate each mixture and visually evaluate the results.
- Score the findings in the same way that you would for RPLA: (full agglutination),,/ or – (absence of agglutination).
- CPE samples do not agglutinate control latex beads. It should be noted that samples having a high concentration of CPE provide negative or moderately positive results, whereas diluted samples produce complete agglutination.
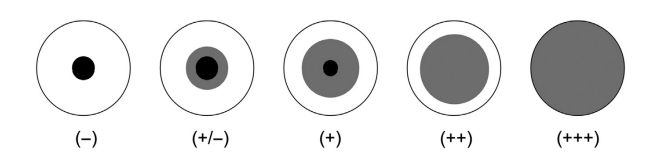
The sensitivity is determined by the purity of the immunoglobulins utilised in the latex bead production. The SLAT sensitivity with purified CPE is 100 ng ml-1 when the immunoglobulin G fraction isolated from rabbit anti-CPE serum is used to sensitise latex beads. However, by utilising particular anti-CPE immunoglobulins isolated by immunoaffinity, a lower limit of detection of 0.1 ng ml-1 can be achieved.
Enzyme-Linked Immunosorbent Assays
Several enzyme-linked immunosorbent assay (ELISA) approaches for CPE titration in various samples, including patient stools, have been proposed. TECHLAB has an ELISA kit available (Blacksburg, VA). A common protocol looks like this:
- Using rabbit anti-CPE immunoglobulins, coat a microtiter plate (100 ml of a 5 mg ml1 solution in PBS). Seal the plate and incubate it overnight at 22 degrees Celsius before washing it four times with PBS containing 0.05% Tween20 (PBST).
- Fill the antibody-coated plate with CPE standard and test samples (100 ml diluted in PBST), then seal it and incubate at 37 C for 90 minutes. Wash the plates as indicated earlier, then incubate for 90 minutes at 37 C in the presence of anti-CPE immunoglobulin G (IgG) horseradish peroxidase conjugate (100 ml diluted in PBST containing 1% normal rabbit serum).
- After washing, add 100 ml of ABTS-H2O2 solution to each well, containing 0.4 mM 2,20 -azino-di(3-ethylbenzothiazoline-6-sulphonate) (ABTS) and 1.3 mM H2O2 in 0.1 mM citrate phosphate buffer, pH 4. Incubate the plate at room temperature for 30 minutes.
- At 403 nm, measure the absorbance. When the absorbance is 0.2 after background adjustment and corresponds to the absorbance in a control noncoated well, the sample is considered to contain CPE. Using pure CPE, estimate the CPE concentration from a standard curve (0–50 ng ml1 ).
The four-layer sandwich ELISA method is a variation:
- Coat 200 ml of goat anti-CPE serum (1-100 dilution in carbonate buffer 0.0015 M Na2CO3 – b 0.035 M NaHCO3, pH 9.6) into each well of an immulon II enzyme immunoassay plate, and incubate the plate overnight at 4 C in a humid room. The plate is then gently shaken on a rotary shaker for 2 minutes after being washed with 100 ml of warmwashing solution containing 0.85% NaCl, 0.05% Tween20, and 0.3% BSA per well. This washing technique should be repeated three times.
- To block the excess binding sites on the microtiter plate, incubate 100 ml of 3% BSA-1% normal goat serum diluted in PBS each well for 30 minutes at 37 C. Then, as indicated above, wash the plate twice.
- In each well, add 100 ml of CPE diluted in 0.05% Tween20 in PBS, and incubate the plates at 37 C for 2 hours. Before repeating the blocking technique described above for 30 minutes at 37 C, wash each well once.
- After washing the plate twice, add 200 ml of rabbit antitoxin diluted 1:200 with 0.85% NaCl, 0.05% Tween20, and 1% BSA to each well and incubate for 2 hours at 37 degrees Celsius. Wash three times, then add 200 ml of conjugate (1:800 dilution of goat antirabbit immunoglobulin G conjugated with alkaline phosphatase in PBS-0.05% Tween20) for 2 hours at 37 C. Add 200 cc of heated substrate (0.1% p-nitrophenol phosphate-10% diethanolamine0.01% MgCl2, pH 9.6) after three additional washes.
- Allow the reaction to proceed at 37 C for 30 minutes before stopping it with 50 cc of 2 M NaOH.
- At 405 nm, read the results spectrophotometrically. Perform the test in duplicate for each sample. Subtract the absorbances (0.02) in negative controls that do not get CPE or sample to get the absorbances. Positive values are those that are more than 0.1.
ELISA has a sensitivity of 1-25 ng ml1 for purified CPE in aqueous solution and 5-500 ng g1 for CPE in faeces samples. The decrease in sensitivity caused by digestion of the IgG used to coat the polystyrene surface is attributed to protease activity in some samples. This can be avoided by adding 1% serum albumin to the samples.
Detection of Enterotoxigenic C. perfringens
- For the identification of enterotoxigenic strains and C. perfringens typing, DNA-based approaches such as PCR, standard PCR, nested PCR, real-time PCR, loop-mediated isothermal amplification, and DNA-DNA hybridization have been developed.
- Multiplex PCR allows for the detection of many toxin genes at the same time. The alpha-toxin gene, which corresponds to a marker of the C. perfringens species and is present in all strains except for a few unusual strains, and the cpe gene, which is diagnostic of the strains involved in food poisoning, have been identified using a duplex PCR.
- The detection level for C. perfringens varies from 103 to 105 cells per gramme of faeces or food sample, and 10 cells per gramme when enrichment culture is utilised.
- The advantage of PCR is that it does not require C. perfringens sporulation and yields accurate findings with culture in standard growth media.
- Among the stool preparation protocols, a quick technique is as follows: A 1 g faeces sample is homogenised in 9 ml of distilled water, centrifuged, and the supernatant is discarded.
- In 0.2 ml of Instagen, the particle is resuspended (BioRad). The mixture is incubated for 30 minutes at 55 degrees Celsius, vortexed rapidly for 10 seconds, and then incubated for 10 minutes at 100 degrees Celsius.
- The mixture is vortexed again for 10 seconds before being centrifuged (10 min at 10 000 rpm). For PCR amplification, supernatant (3 ml) is used, both undiluted and diluted 1:10 in distilled water containing 3% BSA.
Treatment and Prevention
- The clinical symptoms of C. perfringens type A food poisoning are commonly mild and the clinical course is self-limited. Antibiotic treatment is typically not required.
- Rehydration drinks containing important electrolytes are useful for managing the dehydration caused by diarrhea. CPE-producing C. perfringens strains are broadly distributed in the environment, including foods, soil, and kitchen surfaces.
- Therefore, contamination events to food may occur during the cooking process (food ingredients, cooking, and preserving). For this reason, completely protecting against C. perfringens contamination to food(s) is likely to be difficult.
- Fortunately, a large number of CPE-producing bacterial cells are needed to induce C. perfringens food poisoning, which is because many ingested C. perfringens vegetative cells are killed when exposed to the acidity of the stomach.
- Collectively, the most important factors preventing and controlling C. perfringens type A foodborne illness are ‘careful cooking’ and ‘careful storage,’ which prohibit the vegetative growth of C. perfringens in cooked foods.
- Therefore, reducing the total amount or size of food for heating and cooling (it is difficult to archive high internal temperatures in large amounts of food), appropriate storage at less than 10 C for cooked foods before serving (growth rates of C. perfringens vegetative cells rapidly decrease at temperatures below 15 C, with no growth occurring at 6 C), and the immediate consumption of served foods as soon as possible are the best ways to prevent C. perfringens food poisoning.
- Biologically, the growth of C. perfringens strains is affected by water activity (aw), reduction potential (Eh), pH (optimal growth at pH 6 to 7), and chemical preservatives (e.g., NaCl).
- These preservation factors also control the growth of C. perfringens vegetative cells and inhibit the outgrowth of germinating C. perfringens spores in food.
FAQ
What is Clostridium perfringens food poisoning?
Clostridium perfringens food poisoning is a type of bacterial foodborne illness caused by the consumption of food contaminated with the bacterium Clostridium perfringens.
What are the symptoms of Clostridium perfringens food poisoning?
The symptoms of Clostridium perfringens food poisoning typically include abdominal cramps, watery diarrhea, and sometimes nausea. Vomiting and fever are generally absent or mild.
How long does it take for symptoms to appear after consuming contaminated food?
The symptoms of Clostridium perfringens food poisoning usually appear within 6 to 24 hours after consuming contaminated food, with an average onset time of around 8 to 12 hours.
What types of foods are commonly associated with Clostridium perfringens food poisoning?
Clostridium perfringens food poisoning is often linked to foods that are prepared in large quantities and then kept warm for extended periods, such as stews, gravies, and casseroles. Meats, particularly poultry and beef, are frequently implicated.
How does Clostridium perfringens contaminate food?
Clostridium perfringens is commonly found in the intestines of humans and animals, as well as in the environment. It can contaminate food when proper food handling and temperature control practices are not followed, allowing the bacterium to multiply and produce toxins.
Can Clostridium perfringens food poisoning be prevented?
Clostridium perfringens food poisoning can be prevented by practicing good food hygiene and safety measures. This includes proper cooking and reheating of foods, avoiding prolonged temperature abuse, and promptly refrigerating leftovers.
How is Clostridium perfringens food poisoning diagnosed?
Diagnosis of Clostridium perfringens food poisoning is typically based on the presence of characteristic symptoms, along with a history of consuming contaminated food. Laboratory testing can be performed to confirm the presence of the bacterium or its toxins in food samples or fecal specimens.
Is treatment necessary for Clostridium perfringens food poisoning?
In most cases, treatment for Clostridium perfringens food poisoning is not necessary as the illness is self-limiting and resolves within 24 to 48 hours. It is important to stay hydrated by drinking plenty of fluids.
Are antibiotics used to treat Clostridium perfringens food poisoning?
Antibiotics are generally not recommended for the treatment of Clostridium perfringens food poisoning, as they do not significantly alter the course of the illness. However, in severe cases or in individuals with underlying health conditions, antibiotics may be prescribed.
Can Clostridium perfringens food poisoning be fatal?
Clostridium perfringens food poisoning is typically a mild and self-limiting illness. However, in rare cases, complications can arise, especially in individuals with weakened immune systems or other underlying health conditions. Fatalities associated with Clostridium perfringens food poisoning are extremely rare.
References
- Miyamoto, K., & Nagahama, M. (2016). Clostridium: Food Poisoning by Clostridium perfringens. Encyclopedia of Food and Health, 149–154.
- Chukwu, E. E., Nwaokorie, F. O., Coker, A. O., Avila-Campos, M. J., Solis, R. L., Llanco, L. A., & Ogunsola, F. T. (2016). Detection of toxigenic Clostridium perfringens and Clostridium botulinum from food sold in Lagos, Nigeria. Anaerobe, 42, 176–181.
- Popoff, M. R. (2014). CLOSTRIDIUM | Detection of Enterotoxin of Clostridium perfringens. Encyclopedia of Food Microbiology, 474–480 (https://www.sciencedirect.com/science/article/pii/B9780123847300000690)
- Chukwu, E. E., Nwaokorie, F. O., Coker, A. O., Avila-Campos, M. J., Solis, R. L., Llanco, L. A., & Ogunsola, F. T. (2016). Detection of toxigenic Clostridium perfringens and Clostridium botulinum from food sold in Lagos, Nigeria. Anaerobe, 42, 176–181. doi:10.1016/j.anaerobe.2016.10.0
- https://www.cdc.gov/foodsafety/diseases/clostridium-perfringens.html#:~:text=perfringens%20food%20poisoning%20have%20diarrhea,of%20fluids%20to%20prevent%20dehydration.
- https://www.msdmanuals.com/en-in/professional/infectious-diseases/anaerobic-bacteria/clostridium-perfringens-food-poisoning
- https://www.peacehealth.org/medical-topics/id/te6324
- https://myhealth.alberta.ca/Health/pages/conditions.aspx?hwid=te6324
- http://eknygos.lsmuni.lt/springer/655/41-78.pdf
- https://www.foodsafety.gov/blog/prevent-illness-c-perfringens
